

HPC Application Performance on Dell PowerEdge C6620 with INTEL 8480+ SPR
Thu, 09 Nov 2023 15:46:41 -0000
|Read Time: 0 minutes
Overview
With a robust HPC and AI Innovation Lab at the helm, Dell continues to ensure that PowerEdge servers are cutting-edge pioneers in the ever-evolving world of HPC. The latest stride in this journey comes in the form of the Intel Sapphire Rapids processor, a powerhouse of computational prowess. When combined with the cutting-edge infrastructure of the Dell PowerEdge 16th generation servers, a new era of performance and efficiency dawns upon the HPC landscape. This blog post provides comprehensive benchmark assessments spanning various verticals within high-performance computing.
It is Dell Technologies’ goal to help accelerate time to value for customers, as well as leverage benchmark performance and scaling studies to help plan out their environments. By using Dell’s solutions, customers spend less time testing different combinations of CPU, memory, and interconnect, or choosing the CPU with the sweet spot for performance. Additionally, customers do not have to spend time deciding which BIOS features to tweak for best performance and scaling. Dell wants to accelerate the set-up, deployment, and tuning of HPC clusters to enable customers real value while running their applications and solving complex problems (such as weather modeling).
Testbed Configuration
This study conducted benchmarking on high-performance computing applications using Dell PowerEdge 16th generation servers featuring Intel Sapphire Rapids processors.
Benchmark Hardware and Software Configuration
Table 1. Test bed system configuration used for this benchmark study
Platform | Dell PowerEdge C6620 |
Processor | Intel Sapphire Rapids 8480+ |
Cores/Socket | 56 (112 total) |
Base Frequency | 2.0 GHz |
Max Turbo Frequency | 3.80 GHz |
TDP | 350 W |
L3 Cache | 105 MB |
Memory | 512 GB | DDR5 4800 MT/s |
Interconnect | NVIDIA Mellanox ConnectX-7 NDR 200 |
Operating System | Red Hat Enterprise Linux 8.6 |
Linux Kernel | 4.18.0-372.32.1 |
BIOS | 1.0.1 |
OFED | 5.6.2.0.9 |
System Profile | Performance Optimized |
Compiler | Intel OneAPI 2023.0.0 | Compiler 2023.0.0 |
MPI | Intel MPI 2021.8.0 |
Turbo Boost | ON |
Interconnect | Mellanox NDR 200 |
Application | Vertical Domain | Benchmark Datasets |
OpenFOAM | Manufacturing - Computational Fluid Dynamics (CFD) | Motorbike 50 M 34 M and 20 M cell mesh |
Weather Research and Forecasting (WRF) | Weather and Environment | |
Large-scale Atomic/Molecular Massively Parallel Simulator (LAMMPS) | Molecular Dynamics | Rhodo, EAM, Stilliger Weber, tersoff, HECBIOSIM, and Airebo |
GROMACS | Life Sciences – Molecular Dynamics | HECBioSim Benchmarks – 3M Atoms, Water, and Prace LignoCellulose |
CP2K | Life Sciences | H2O-DFT-LS-NREP- 4, 6 H2O-64-RI-MP 2 |
Performance Scalability for HPC Application Domain
Vertical – Manufacturing | Application - OPENFOAM
OpenFOAM is an open-source computational fluid dynamics (CFD) software package renowned for its versatility in simulating fluid flows, turbulence, and heat transfer. It offers a robust framework for engineers and scientists to model complex fluid dynamics problems and conduct simulations with customizable features. This study worked on OpenFOAM version 9, which have been compiled with Intel ONE API 2023.0.0 and Intel MPI 2021.8.0 compilers. For successful compilation and optimization with the Intel compilers, additional flags such as '-O3 -xSAPPHIRERAPIDS -m64 -fPIC' have been added.
The tutorial case under the simpleFoam solver category, motorbike, were used for evaluating the performance of the OpenFOAM package on intel 8480+ processors. Three different types of grids were generated such as 20 M, 34 M, and 50 M cells using the blockMesh and snappyHexMesh utilities of OpenFOAM. Each run was conducted with full cores (112 cores per node) and from a single node to sixteen nodes, while scalability tests were done for all three sets of grids. The steady state simpleFoam solver execution time was noted as performance numbers.
The figure below shows the application performance for all the datasets:
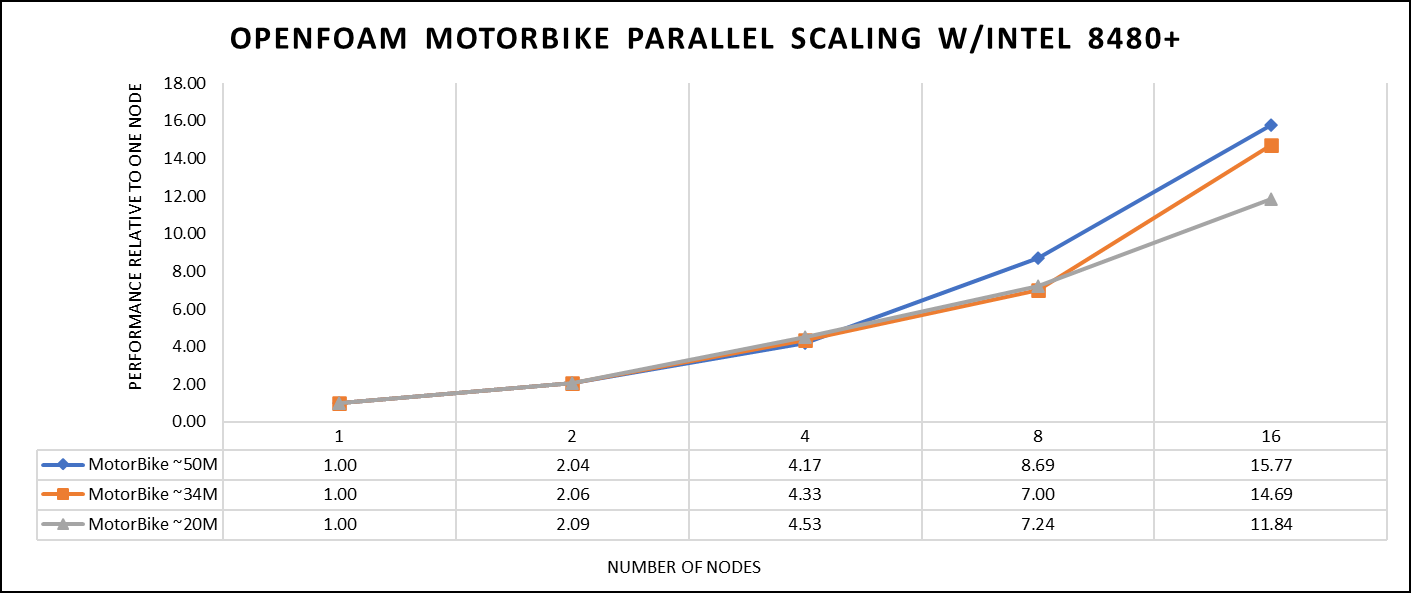
Figure 1. The scaling performance of the OpenFOAM Motorbike dataset using the Intel 8480+ processor, with a focus on performance compared to a single node.
The results are non-dimensionalized with single node result, with the scalability depicted in Figure 1. The Intel-compiled binaries of OpenFOAM shows linear scaling from a single node to sixteen nodes on 8480+ processors for higher dataset (50 M). For other datasets with 20 M and 34 M cells, the linear scaling was shown up to eight nodes and from eight nodes to sixteen nodes the scalability was reduced.
Achieving satisfactory results with smaller datasets can be accomplished using fewer processors and nodes. Nonetheless, augmenting the node count; therefore, the processor count, in relation to the solver's computation time, leads to increased inter-processor communication, later extending the overall runtime. Consequently, higher node counts prove more beneficial when handling larger datasets within OpenFOAM simulations.
Vertical – Weather and Environment | Application - WRF
The Weather Research and Forecasting model (WRF) is at the forefront of meteorological and atmospheric research, with its latest version being a testament to advancements in high-performance computing (HPC). WRF enables scientists and meteorologists to simulate and forecast complex weather patterns with unparalleled precision. This study involved working on WRF version 4.5, which have been compiled with Intel ONEAPI 2023.0.0 and Intel MPI 2021.8.0 compilers. For successful compilation and optimization with the Intel compilers, additional flags such as ' -O3 qopt-zmm-usage=high –xSAPPHIRERAPIDS -fpic’ were used.
The dataset used in this study is CONUS v4.4, meaning the model's grid, parameters, and input data are set up to focus on weather conditions within the continental United States. This configuration is particularly useful for researchers, meteorologists, and government agencies who need high-resolution weather forecasts and simulations tailored to this specific geographic area. The configuration details, such as grid resolution, atmospheric physics schemes, and input data sources, can vary depending on the specific version of WRF and the goals of the modeling project. This study predominantly adhered to the default input configuration, making minimal alterations or adjustments to the source code or input file. Each run was conducted with full cores (112 cores per node). The scalability tests were done from a single node to sixteen nodes, and the performance metric in “sec” was noted.
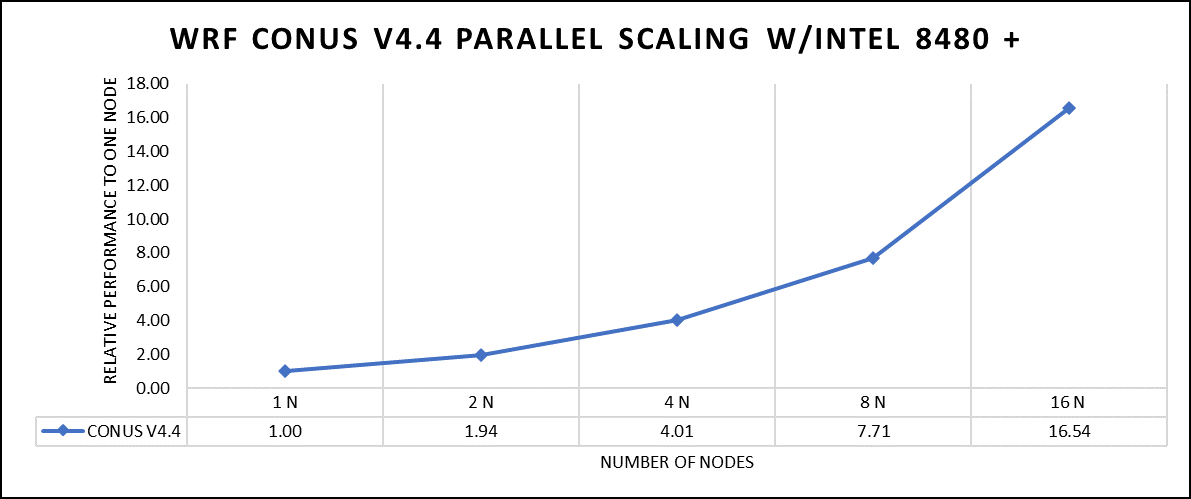
Figure 2. The scaling performance of the WRF CONUS dataset using the Intel 8480+ processor, with a focus on performance compared to a single node.
The INTEL compiled binaries of WRF show linear scaling from a single node to sixteen nodes on 8480+ processors for the new CONUS v4.4. For the best performance with WRF, the impact of the tile size, process, and threads per process should be carefully considered. Given that the memory and DRAM bandwidth constrain the application, the team opted for the latest DDR5 4800 MT/s DRAM for test evaluations. Additionally, it is crucial to consider the BIOS settings, particularly SubNUMA configurations, as these settings can significantly influence the performance of memory-bound applications, potentially leading to improvements ranging from one to five percent.
For more detailed BIOS tuning recommendations, see the previous blog post on optimizing BIOS settings for optimal performance.
Vertical – Molecular Dynamics | Application – LAMMPS
LAMMPS, which stands for Large-scale Atomic/Molecular Massively Parallel Simulator, is a powerful tool for HPC. It is specifically designed to harness the immense computational capabilities of HPC clusters and supercomputers. LAMMPS allows researchers and scientists to conduct large-scale molecular dynamics simulations with remarkable efficiency and scalability. This study worked on LAMMPS, the 15 June 2023 version, which have been compiled with Intel ONEAPI 2023.0.0 and Intel MPI 2021.8.0 compilers. For successful compilation and optimization with the Intel compilers, additional flags such as “ -O3 qopt-zmm-usage=high –xSAPPHIRERAPIDS -fpic,” were used.
The team opted for the default INTEL package, which offers optimized atom pair styles for vector instructions on Intel processors. The team also tried running some benchmarks which are not supported with the INTEL package to check the performance and scaling. The performance metric for this benchmark is nanoseconds per day where higher is considered better.
There are two factors that were considered when compiling data for comparison: the number of nodes and the core count. Below are the results of performance improvement observed on processor 8480+ with 112 cores:
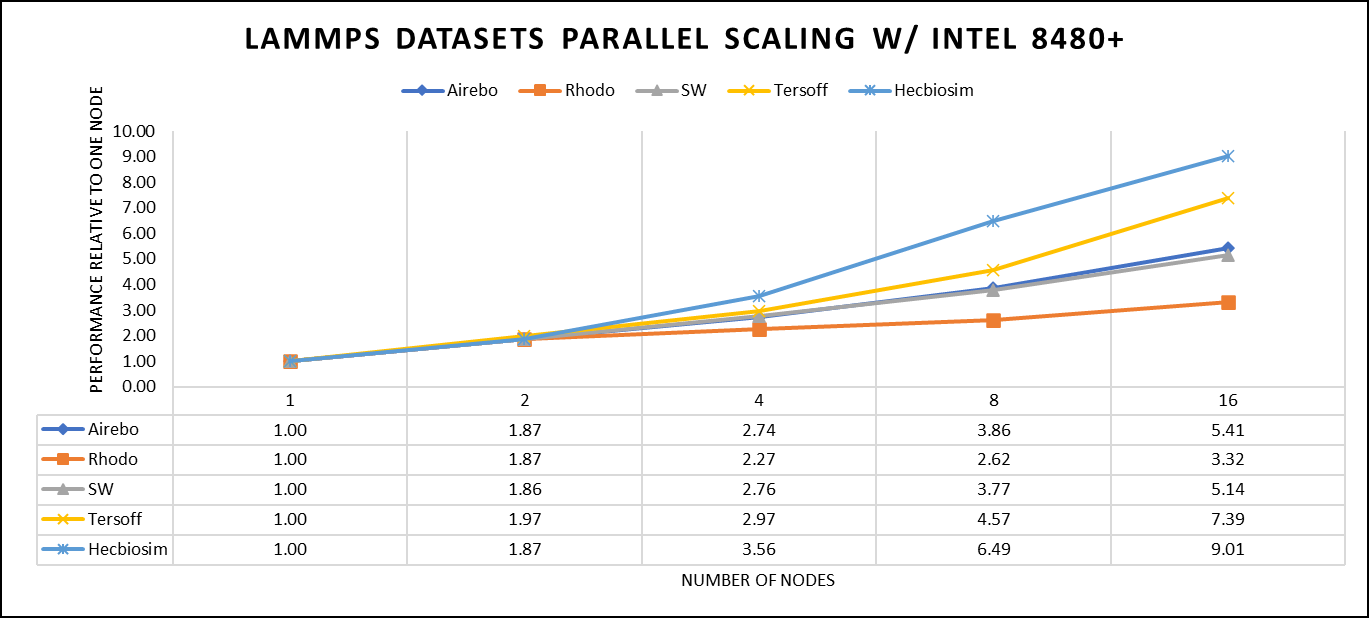
Figure 3. The scaling performance of the LAMMPS datasets using the Intel 8480+ processor, with a focus on performance compared to a single node.
Figure 3 shows the scaling of different LAMMPS datasets. Noticeable enhancement in scalability is evident with the increment in atom size and step size. The examination involved two datasets, EAM and Hecbiosim, each containing over 3 million atoms. The results indicated better scalability when compared to the other datasets analyzed.
Vertical – Molecular Dynamics | Application - GROMACS
GROMACS, a high-performance molecular dynamics software, is a vital tool for HPC environments. Tailored for HPC clusters and supercomputers, GROMACS specializes in simulating the intricate movements and interactions of atoms and molecules. Researchers in diverse fields, including biochemistry and biophysics, rely on its efficiency and scalability to explore complex molecular processes. GROMACS is used for its ability to harness the immense computational power of HPC, allowing scientists to conduct intricate simulations that reveal critical insights into atomicatomic-level behaviours, from biomolecules to chemical reactions and materials. This study worked on GROMACS 2023.1 version, which has been compiled with Intel ONEAPI 2023.0.0 and Intel MPI 2021.8.0 compilers. For successful compilation and optimization with the Intel compilers, additional flags such as “ -O3 qopt-zmm-usage=high –xSAPPHIRERAPIDS -fpic,” were used.
The team curated a range of datasets for the benchmarking assessments. First, the team included "water GMX_50 1536" and "water GMX_50 3072," which represent simulations involving water molecules. These simulations are pivotal for gaining insights into solvation, diffusion, and the water's behavior in diverse conditions. Next, the team incorporated "HECBIOSIM 14 K" and "HECBIOSIM 30 K" datasets, which were specifically chosen for their ability to investigate intricate systems and larger biomolecular structures. Lastly, the team included the "PRACE Lignocellulose" dataset, which aligns with the benchmarking objectives, particularly in the context of lignocellulose research. These datasets collectively offer a diverse array of scenarios for the benchmarking assessments.
The performance assessment was based on the measurement of nanoseconds per day (ns/day) for each dataset, providing valuable insights into the computational efficiency. Additionally, the team paid careful attention to optimizing the mdrun tuning parameters (i.e, ntomp, dlb tunepme nsteps, etc )in every test run to ensure accurate and reliable results. The team examined the scalability by conducting tests spanning from a single node to sixteen nodes.
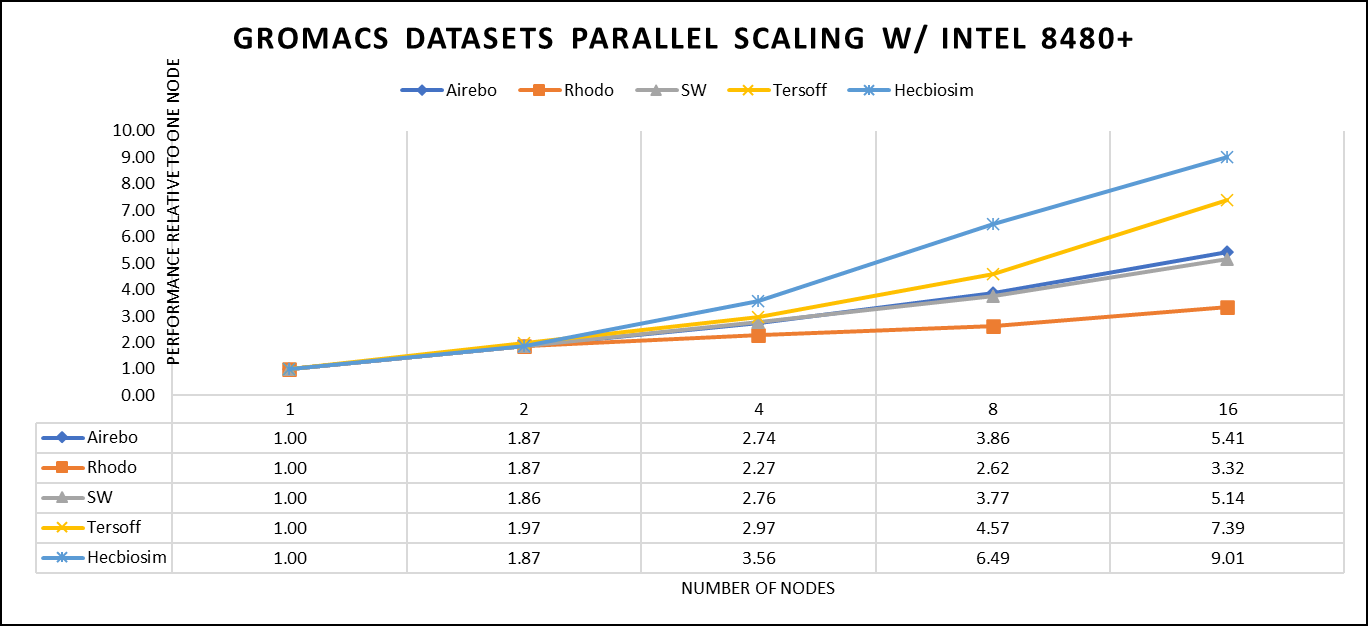
Figure 4. The scaling performance of the GROMACS datasets using the Intel 8480+ processor, with a focus on performance compared to a single node.
For ease of comparison across the various datasets, the relative performance has been included into a single graph. However, each dataset behaves individually when performance is considered, as each uses different molecular topology input files (tpr), and configuration files.
The team achieved the expected linear performance scalability for GROMACS of up to eight nodes All cores in each server were used while running these benchmarks. The performance increases are close to linear across all the dataset types; however, there is a drop in the larger number of nodes due to the smaller dataset size and the simulation iterations.
Vertical – Molecular Dynamics | Application – CP2K
CP2K is a versatile computational software package that covers a wide range of quantum chemistry and solid-state physics simulations, including molecular dynamics. It is not strictly limited to molecular dynamics but is instead a comprehensive tool for various computational chemistry and materials science tasks. While CP2K is widely used for molecular dynamics simulations, it can also perform tasks like electronic structure calculations, ab initio molecular dynamics, hybrid quantum mechanics/molecular mechanics (QM/MM) simulations, and more.
This study worked on the CP2K 2023.1 version, which has been compiled with Intel ONEAPI 2023.0.0 and Intel MPI 2021.8.0 compilers. For successful compilation and optimization with the Intel compilers, additional flags such as “ -O3 qopt-zmm-usage=high –xSAPPHIRERAPIDS -fpic,” were used.
Focusing on high-performance computing (HPC), the team used specific datasets optimized for computational efficiency. The first dataset, "H2O-DFT-LS-NREP-4,6," was configured for HPC simulations and calculations, emphasizing the modeling of water (H2O) using Density Functional Theory (DFT). The appended "NREP-4,6" parameter settings were fine-tuned to ensure efficient HPC performance. The second dataset, "H2O-64-RI-MP2," was exclusively crafted for HPC applications and revolved around the examination of a system consisting of 64 water molecules (H2O). By employing the Resolution of Identity (RI) method with the Møller–Plesset perturbation theory of second order (MP2), this dataset demonstrated the significant computational capabilities of HPC for conducting advanced electronic structure calculations within a high-molecule-count environment. The team examined the scalability by conducting tests spanning from a single node to sixteen nodes.
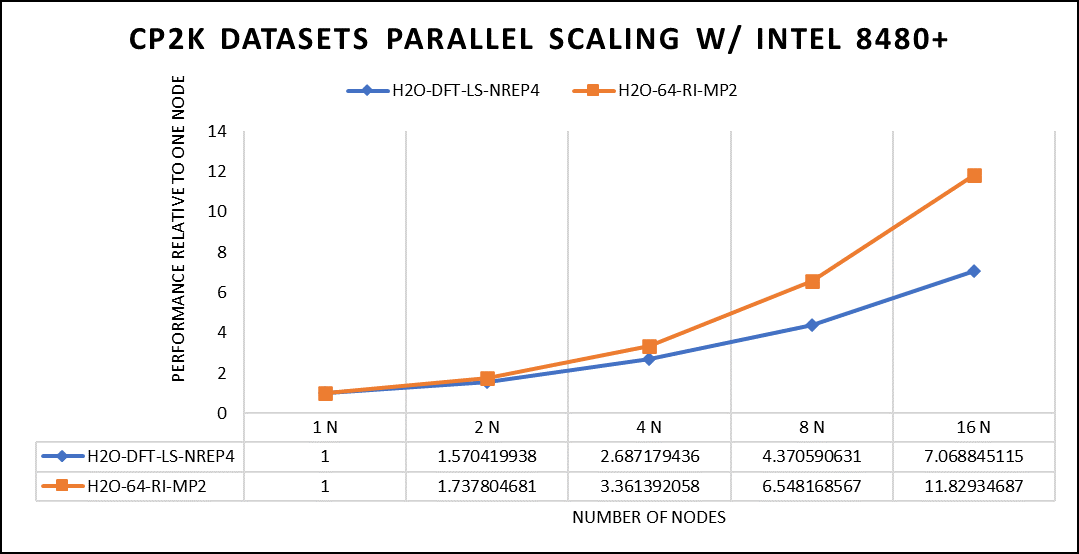
Figure 5. The scaling performance of the CP2K datasets using the Intel 8480+ processor, with a focus on performance compared to a single node.
The datasets represent a single-point energy calculation employing linear-scaling Density Functional Theory (DFT). The system consists of 6144 atoms confined within a 39 Å^3 simulation box, which translates to 2048 water molecules. To adjust the computational workload, you can modify the NREP parameter within the input file.
Performing with NREP6 necessitates more than 512 GB of memory on a single node. Failing to meet this requirement will result in a segmentation fault error. These benchmarking efforts encompass configurations involving up to 16 computational nodes. Optimal performance is achieved when using NREP4 and NREP6 in Hybrid mode, which combines Message Passing Interface (MPI) and Open Multi-Processing (OpenMP). This configuration exhibits the best scaling performance, particularly on four to eight nodes. However, it is worth noting that scaling beyond eight nodes does not exhibit a strictly linear performance improvement. Figure 5 depicts the outcomes when using Pure MPI, using 112 cores with a single thread per core.
Conclusion
With equivalent core counts, the prior generation of Intel Xeon processors can match the performance of the Sapphire Rapids counterpart. However, achieving this level of performance necessitates doubling the number of nodes. Therefore, a single 350W node equipped with the 8480+ processor can deliver comparable performance when compared to using two 500W nodes with the 8358 processor. In addition to optimizing the BIOS settings as outlined in our INTEL-focused blog, the team advises disabling Hyper-threading specifically for the benchmarks discussed in this article. However, for different types of workloads, the team recommends conducting thorough testing and enabling Hyper-threading if it proves beneficial. Furthermore, for this performance study, the team highly recommends using the Mellanox NDR 200 interconnect.
Related Blog Posts

HPC Application Performance on Dell PowerEdge R6625 with AMD EPYC- GENOA
Wed, 08 Nov 2023 21:09:35 -0000
|Read Time: 0 minutes
The AMD EPYC 9354 Processor, when integrated into the Dell R6625 server, offers a formidable solution for high-performance computing (HPC) applications. Genoa, which is built on the Zen 4 architecture, delivers exceptional processing power and efficiency, making it a compelling choice for demanding HPC workloads. When paired with the PowerEdge R6625's robust infrastructure and scalability features, this CPU enhances server performance, enabling efficient and reliable execution of HPC applications. These features make it an ideal choice for HPC application studies and research.
At Dell Technologies, it’s our goal to help accelerate time to value for our customers. Dell wants to help customers leverage our benchmark performance and scaling studies to help plan out their environments. By utilizing our expertise, customers don’t have to spend time testing different combinations of CPU, memory and interconnect or choosing the CPU with the sweet spot for performance. This also saves time, as customers don’t have to spend time deciding which BIOS features to tweak for best performance and scaling. Dell wants to accelerate the set-up, deployment, and tuning of HPC clusters to enable customers to get the real value- running their applications and solving complex problems like manufacturing better products for their customers.
Testbed configuration
Benchmarking for high-performance computing applications was carried out using Dell PowerEdge 16G servers equipped with AMD EPYC 9354 32-Core Processor.
Table 1. Test bed system configuration used for this benchmark study
Platform | Dell PowerEdge R6625 |
Processor | AMD EPYC 9354 32-Core Processor |
Cores/Socket | 32 (64 total) |
Base Frequency | 3.25 GHz |
Max Turbo Frequency | 3.75 GHz |
TDP | 280 W |
L3 Cache | 256 MB |
Memory | 768 GB | DDR5 4800 MT/s |
Interconnect | NVIDIA Mellanox ConnectX-7 NDR 200 |
Operating System | RHEL 8.6 |
Linux Kernel | 4.18.0-372.32.1 |
BIOS | 1.0.1 |
OFED | 5.6.2.0.9 |
System Profile | Performance Optimized |
Compiler | AOCC 4.0.0 |
MPI | OpenMPI 4.1.4 |
Turbo Boost | ON |
Interconnect | Mellanox NDR 200 |
Application | Vertical Domain | Benchmark Datasets |
OpenFOAM | Manufacturing - Computational Fluid Dynamics (CFD) | Motorbike 50M 34M and 20M cell mesh |
Weather Research and Forecasting (WRF) | Weather and Environment |
|
Large-scale Atomic/Molecular Massively Parallel Simulator (LAMMPS) | Molecular Dynamics | Rhodo, EAM, Stilliger Weber, tersoff, HECBIOSIM, and Airebo |
GROMACS | Life Sciences – Molecular Dynamics | HECBioSim Benchmarks – 3M Atoms , Water and Prace LignoCellulose |
CP2K | Life Sciences | H2O-DFT-LS-NREP- 4,6 H2O-64-RI-MP 2 |
Performance Scalability for HPC Application Domain
Vertical – Manufacturing | Application - OPENFOAM
OpenFOAM is an open-source computational fluid dynamics (CFD) software package renowned for its versatility in simulating fluid flows, turbulence, and heat transfer. It offers a robust framework for engineers and scientists to model complex fluid dynamics problems and conduct simulations with customizable features. In this study, worked on OpenFOAM version 9, which have been compiled with gcc 11.2.0 with OPENMPI 4.1.5. For successful compilation and optimization on the AMD EPYC processors, additional flags such as ' -O3 -znver4' have been added.
The tutorial case under the simpleFoam solver category, motorBike, has been used to evaluate the performance of OpenFOAM package on AMD EPYC 9354 processors. Three different types of grids were generated such as 20M, 34M, and 50M cells using the blockMesh and snappyHexMesh utilities of OpenFOAM. Each run was conducted with full cores (64 cores per node) and from single node to sixteen nodes, The scalability tests were done for all the three sets of grids. The steady state simpleFoam solver execution time was noted down as performance numbers. The figure below shows the application performance for all the datasets.
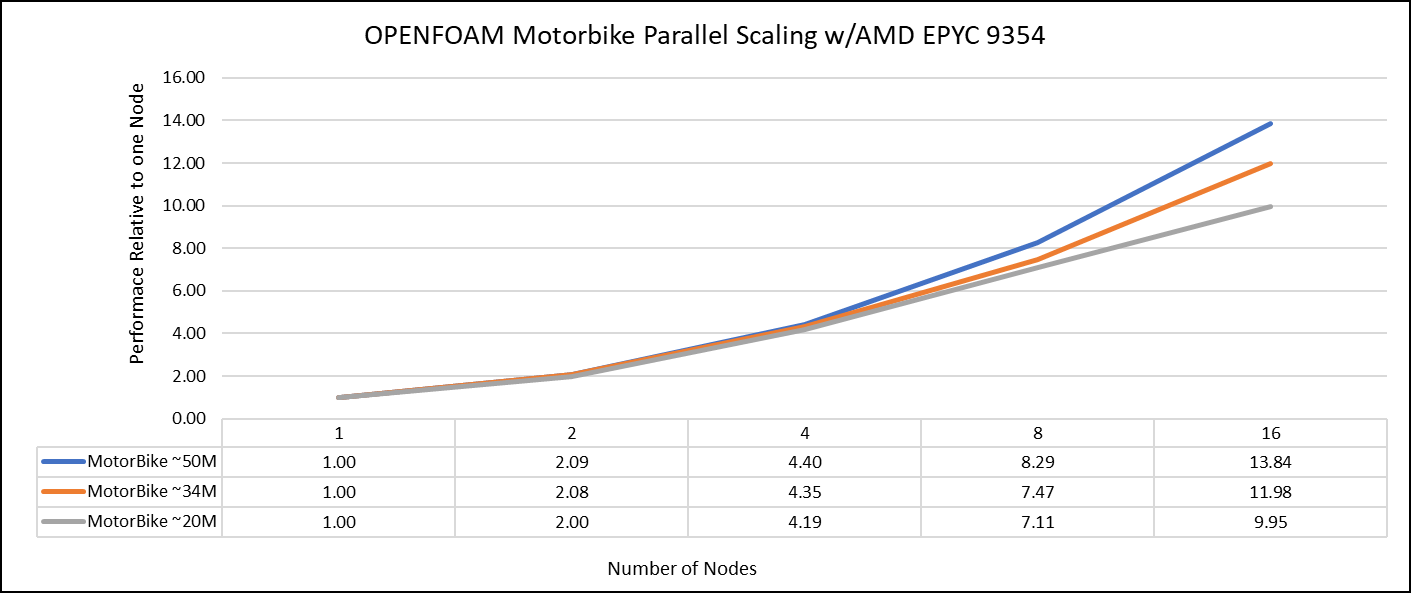
Figure 1: The scaling performance of the OpenFOAM Motorbike dataset using the AMD EPYC Processor, with a focus on performance compared to a single node.
The results are non-dimensionalized with single node result. The scalability is depicted in Figure 1. The OpenFOAM application shows linear scaling from a single node to eight nodes on 9354 processors for higher dataset (50M). For other smaller datasets with 20M and 34M cells, the linear scaling was shown up to four nodes and slightly scaling reduced on eight nodes. For all the datasets (20M, 34M and 50M) on sixteen nodes the scalability was reduced.
Achieving satisfactory results with smaller datasets can be accomplished using fewer processors and nodes, because smaller datasets do not require a higher number of processors. Nonetheless, augmenting the node count, and therefore, the processor count, in relation to the solver's computation time leads to increased interprocessor communication, subsequently extending the overall runtime. Consequently, higher node counts are more beneficial when handling larger datasets within OpenFOAM simulations.
Vertical – Weather and Environment | Application - WRF
The Weather Research and Forecasting model (WRF) is at the forefront of meteorological and atmospheric research, with its latest version being a testament to advancements in high-performance computing (HPC). WRF enables scientists and meteorologists to simulate and forecast complex weather patterns with unparalleled precision. In this study, we have worked on WRF version 4.5, which have been compiled with AOCC 4.0.0 with OPENMPI 4.1.4. For successful compilation and optimization with the AMD EPYC compilers, additional flags such as ' -O3 -znver4' have been added.
The dataset used in our study is CONUS v4.4. This means that the model's grid, parameters, and input data are set up to focus on weather conditions within the continental United States. This configuration is particularly useful for researchers, meteorologists, and government agencies who need high-resolution weather forecasts and simulations tailored to this specific geographic area. The configuration details, such as grid resolution, atmospheric physics schemes, and input data sources, can vary depending on the specific version of WRF and the goals of the modeling project. In this study, we have predominantly adhered to the default input configuration, making minimal alterations or adjustments to the source code or input file. Each run was conducted with full cores (64 cores per node) and from single node to sixteen nodes. The scalability tests were conducted and the performance metric in “sec” was noted.
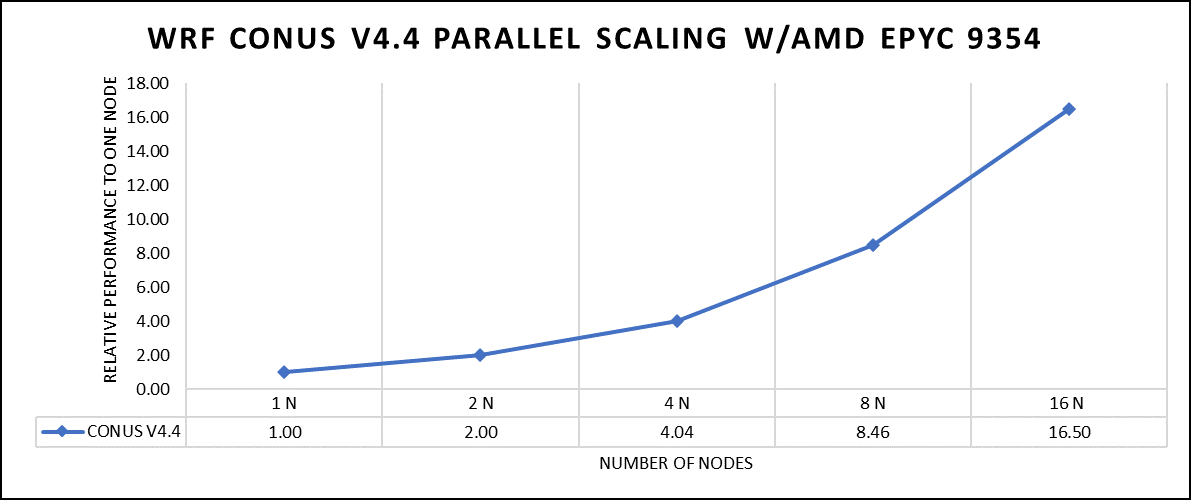
Figure 2: The scaling performance of the WRF CONUS dataset using the AMD EPYC 9354 processor, with a focus on performance compared to a single node.
The AOCC compiled binaries of WRF show linear scaling from a single node to sixteen nodes on 9354 processors for the new CONUS v4.4. For the best performance with WRF, the impact of the tile size, process, and threads per process should be carefully considered. Given that the application is constrained by memory and DRAM bandwidth, we have opted for the latest DDR5 4800 MT/s DRAM for our test evaluations. It is also crucial to consider the BIOS settings, particularly SubNUMA configurations, as these settings can significantly influence the performance of memory-bound applications, potentially leading to improvements ranging from one to five percent. For more detailed BIOS tuning recommendations, please see our previous blog post on optimizing BIOS settings for optimal performance.
Vertical – Molecular Dynamics | Application - LAMMPS
Large-scale Atomic/Molecular Massively Parallel Simulator (LAMMPS) is a powerful tool for HPC. It is specifically designed to harness the immense computational capabilities of HPC clusters and supercomputers. LAMMPS allows researchers and scientists to conduct large-scale molecular dynamics simulations with remarkable efficiency and scalability. In this study, we have worked on LAMMPS, 15 June 2023 version, which have been compiled with AOCC 4.0.0 with OPENMPI 4.1.4. For successful compilation and optimization with the AMD EPYC compilers, additional flags such as ' -O3 -znver4' have been added.
We opted for the non-default package, which offers optimized atom pair styles. We have also tried running some benchmarks which are not supported with default package to check the performance and scaling. Our performance metric for this benchmark is nanoseconds per day, where higher nanoseconds per day is considered a better result .
There are two factors that were considered when compiling data for comparison, the number of nodes and the core count. Figure 3 shows results of performance improvement observed on processor 9354 with 64 cores.
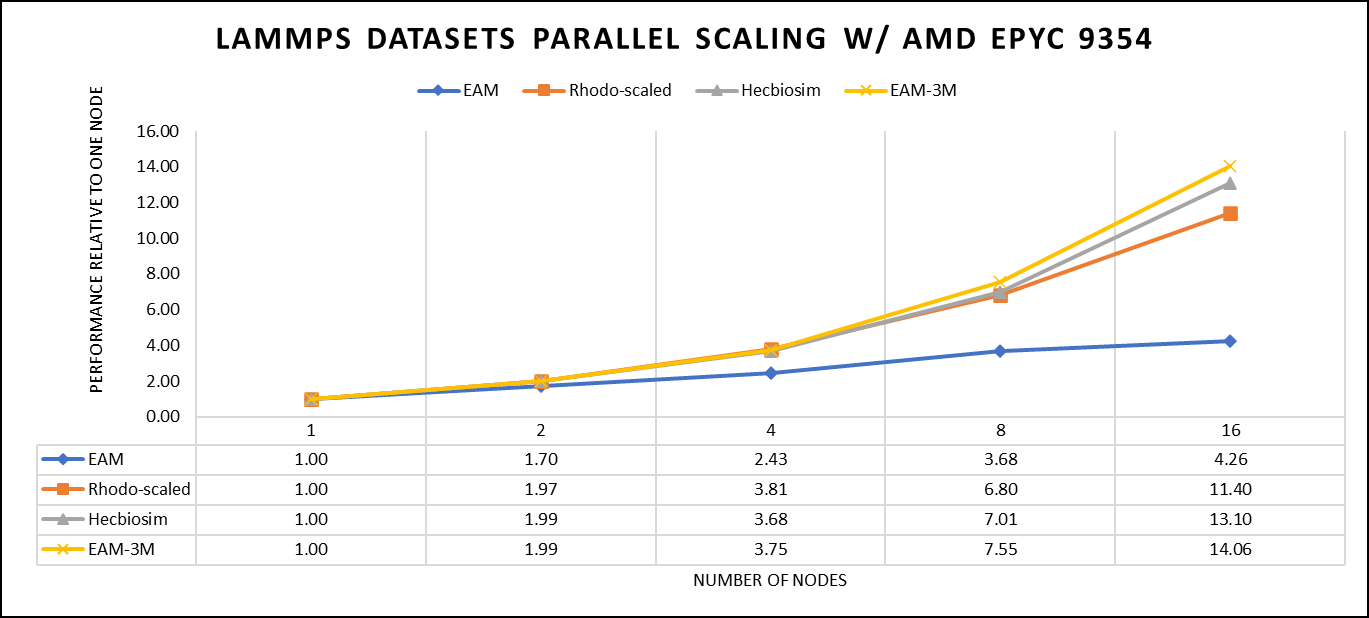
Figure 3: The scaling performance of the LAMMPS datasets using the AMD EPYC 9354 processor, with a focus on performance compared to a single node.
Figure 3 shows the scaling of different LAMMPS datasets. We see a significant improvement in scaling as we increased the atom size and step size. We have tested two datasets EAM and Hecbiosim with more than 3 million atoms and observed a better scalability as compared to other datasets.
Vertical – Molecular Dynamics | Application - GROMACS
GROMACS, a high-performance molecular dynamics software, is a vital tool for HPC environments. Tailored for HPC clusters and supercomputers, GROMACS specializes in simulating the intricate movements and interactions of atoms and molecules. Researchers in diverse fields, including biochemistry and biophysics, rely on its efficiency and scalability to explore complex molecular processes. GROMACS is used for its ability to harness the immense computational power of HPC, allowing scientists to conduct intricate simulations that unveil critical insights into atomic-level behaviors, from biomolecules to chemical reactions and materials. In this study, we have worked on GROMACS 2023.1 version, which have been compiled with AOCC 4.0.0 with OPENMPI 4.1.4. For successful compilation and optimization with the AMD EPYC compilers, additional flags such as ' -O3 -znver4' have been added.
We've curated a range of datasets for our benchmarking assessments. First, we included "water GMX_50 1536" and "water GMX_50 3072," which represent simulations involving water molecules. These simulations are pivotal for gaining insights into solvation, diffusion, and water's behaviour in diverse conditions. Next, we incorporated "HECBIOSIM 14K" and "HECBIOSIM 30K" datasets, which were specifically chosen for their ability to investigate intricate systems and larger biomolecular structures. Lastly, we included the "PRACE Lignocellulose" dataset, which aligns with our benchmarking objectives, particularly in the context of lignocellulose research. These datasets collectively offer a diverse array of scenarios for our benchmarking assessments.
Our performance assessment was based on the measurement of nanoseconds per day (ns/day) for each dataset, providing valuable insights into the computational efficiency. Additionally, we paid careful attention to optimizing the mdrun tuning parameters (i.e, ntomp, dlb tunepme nsteps etc )in every test run to ensure accurate and reliable results. We examined the scalability by conducting tests spanning from a single node to a total of sixteen nodes.
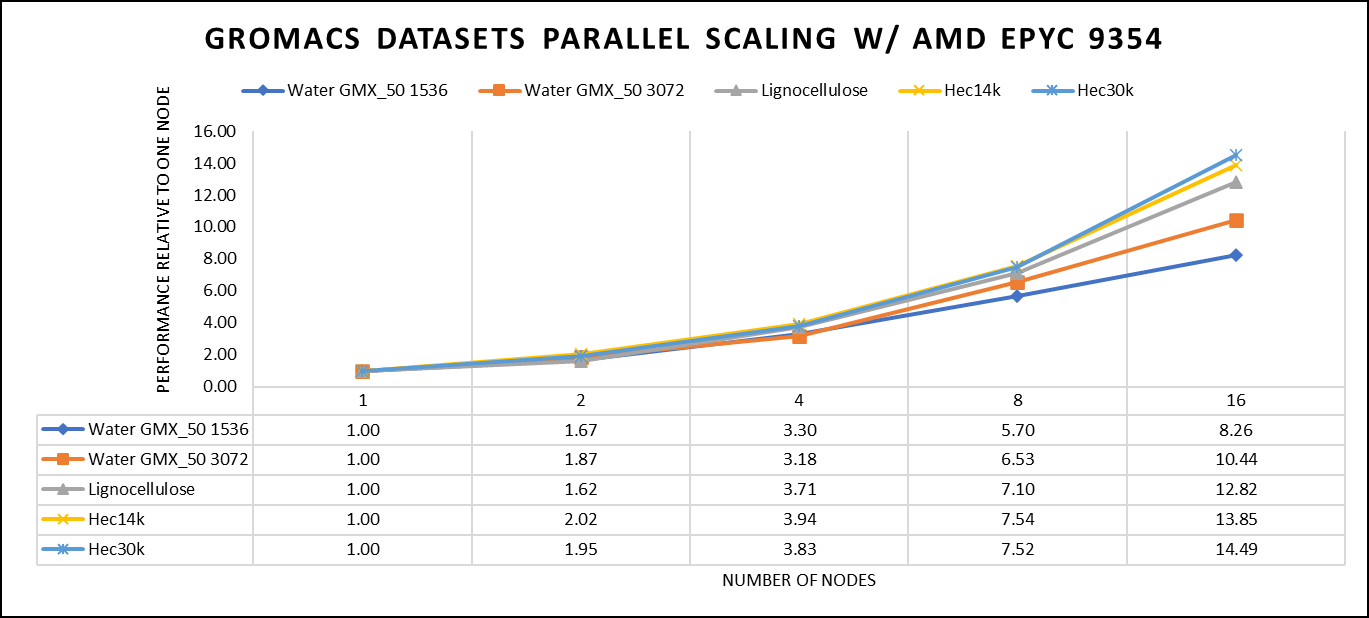
Figure 4: The scaling performance of the GROMACS datasets using the AMD EPYC 9354 processor, with a focus on performance compared to a single node.
For ease of comparison across the various datasets, the relative performance has been included into a single graph. However, it is worth noting that each dataset behaves individually when performance is considered, as each uses different molecular topology input files (tpr), and configuration files.
We were able to achieve the expected performance scalability for GROMACS of up to eight nodes for larger datasets. All cores in each server were used while running these benchmarks. The performance increases are close to linear across all the dataset types, however there is drop in larger number of nodes due to the smaller dataset size and the simulation iterations.
Vertical – Molecular Dynamics | Application – CP2K
CP2K is a versatile computational software package that covers a wide range of quantum chemistry and solid-state physics simulations, including molecular dynamics. It's not strictly limited to molecular dynamics but is instead a comprehensive tool for various computational chemistry and materials science tasks. While CP2K is widely used for molecular dynamics simulations, it can also perform tasks like electronic structure calculations, ab initio molecular dynamics, hybrid quantum mechanics/molecular mechanics (QM/MM) simulations, and more. In this study, we have worked on CP2K 2023.1 version, which have been compiled with AOCC 4.0.0 with OPENMPI 4.1.4. For successful compilation and optimization with the AMD EPYC compilers, additional flags such as ' -O3 -znver4' have been added.
In our study focusing on high-performance computing (HPC), we utilized specific datasets optimized for computational efficiency. The first dataset, "H2O-DFT-LS-NREP-4,6," was configured for HPC simulations and calculations, emphasizing the modeling of water (H2O) using Density Functional Theory (DFT). The appended "NREP-4,6" parameter settings were fine-tuned to ensure efficient HPC performance. The second dataset, "H2O-64-RI-MP2," was exclusively crafted for HPC applications and revolved around the examination of a system comprising 64 water molecules (H2O). By employing the Resolution of Identity (RI) method in conjunction with the Møller–Plesset perturbation theory of second order (MP2), this dataset demonstrated the significant computational capabilities of HPC for conducting advanced electronic structure calculations within a high-molecule-count environment. We examined the scalability by conducting tests spanning from a single node to a total of sixteen nodes.
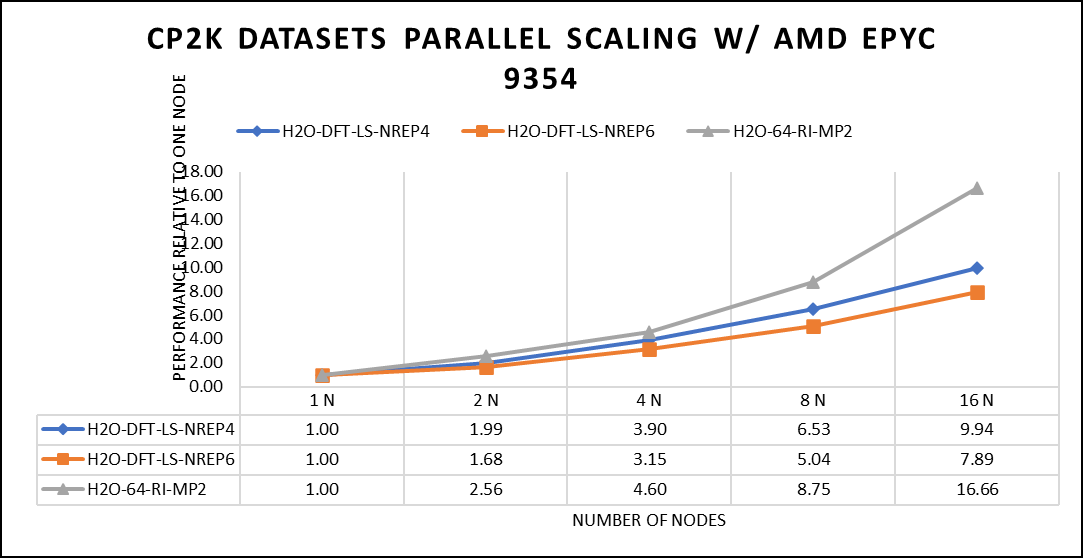
Figure 5: The scaling performance of the CP2K datasets using the AMD EPYC 9354 processor, with a focus on performance compared to a single node.
The datasets represent a single-point energy calculation employing linear-scaling Density Functional Theory (DFT). The system comprises 6144 atoms confined within a 39 Å^3 simulation box, which translates to a total of 2048 water molecules. To adjust the computational workload, you can modify the NREP parameter within the input file.
Our benchmarking efforts encompass configurations involving up to 16 computational nodes .Optimal performance is achieved when using NREP4 and NREP6 in Hybrid mode, which combines MPI (Message Passing Interface) and OpenMP (Open Multi-Processing). This configuration exhibits the best scaling performance, particularly on 4 to 8 nodes. However, it's worth noting that scaling beyond 8 nodes does not exhibit a strictly linear performance improvement. Above figure 5 depict outcomes when using Pure MPI, utilizing 64 cores with a single thread per core.
Conclusion
When considering CPUs with equivalent core counts, the earlier AMD EPYC processors can deliver performance levels like their Genoa counterparts. However, achieving this performance parity may require doubling the number of nodes. To further enhance performance using AMD EPYC processors, we suggest optimizing the BIOS settings as outlined in our previous blog post and specifically disabling Hyper-threading for the benchmarks discussed in this article. various workloads, we recommend conducting comprehensive testing and, if beneficial, enabling Hyper-threading. Additionally, for this performance study, we highly endorse the utilization of the Mellanox NDR 200 interconnect for optimal results.

16G PowerEdge Platform BIOS Characterization for HPC with Intel Sapphire Rapids
Fri, 30 Jun 2023 13:44:52 -0000
|Read Time: 0 minutes
Dell added over a dozen next-generation systems to the extensive portfolio of Dell PowerEdge 16G servers. These new systems are to accelerate performance and reliability for powerful computing across core data centers, large-scale public clouds, and edge locations.
The new PowerEdge servers feature rack, tower, and multi-node form factors, supporting the new 4th-gen Intel Xeon Scalable processors (formerly codenamed Sapphire Rapids). Sapphire Rapids still supports the AVX 512 SIMD instructions, which allow for 32 DP FLOP/cycle. The upgraded Ultra Path Interconnect (UPI) Link speed of 16 GT/s is expected to improve data movement between the sockets. In addition to core count and frequency, Sapphire Rapids-based Dell PowerEdge servers support DDR5 – 4800 MT/s RDIMMS with eight memory channels per processor, which is expected to improve the performance of memory bandwidth-bound applications.
This blog provides synthetic benchmark results and recommended BIOS settings for the Sapphire Rapids-based Dell PowerEdge Server processors. This document contains guidelines that allow the customer to optimize their application for best energy efficiency and provides memory configuration and BIOS setting recommendations for the best out-of-the-box performance and scalability on the 4th Generation of Intel® Xeon® Scalable processor families.
Test bed hardware and software details
Table 1 and Table 2 show the test bed hardware details and synthetic application details. There were 15 BIOS options explored through application performance testing. These options can be set and unset via the Remote Access Control Admin (RACADM) command in Linux or directly when the machines are in the BIOS mode.
Use the following command to set the “HPC Profile” to get the best synthetic benchmark results.
racadm set bios.sysprofilesettings.WorkloadProfile HpcProfile && sudo racadm jobqueue create BIOS.Setup.1-1 -r pwrcycle -s TIME_NOW -e TIME_NA
Once the system is up, use the below command to verify if the setting is enabled.
racadm bios.sysprofilesettings.WorkloadProfile
It should show workload profile set as HPCProfile. Please note that any changes made in BIOS settings on top of the “HPCProfile” will set this parameter to “Not Configured”, while keeping the other settings of “HPCProfile” intact.
Table 1. System details
Component | Dell PowerEdge R660 server (Air cooled) | Dell PowerEdge R760 server (Air cooled) | Dell PowerEdge C-Series (C6620) server (Direct Liquid Cooled) |
SKU | 8452Y | 6430 | 8480+ |
Cores/Socket | 36 | 32 | 56 |
Base Frequency | 2 | 1.9 | 2 |
TDP | 300 | 270 | 350 |
L3Cache | 69.12 MB | 61.44 MB | 10.75 MB |
Operating System | RHEL 8.6 | RHEL 8.6 | RHEL 8.6 |
Memory | 1024 - 64 x 16 | 1024 - 64 x 16 | 512 -32 x 16 |
BIOS | 1.0.1 | 1.0.1 | 1.0.1 |
CPLD | 1.0.1 | 1.0.1 | 1.0.1 |
Interconnect | NDR 400 | NDR 400 | NDR 400 |
Compiler | OneAPI 2023 | OneAPI 2023 | OneAPI 2023 |
Table 2. Synthetic benchmark applications details
Application Name | Version |
High-Performance Linpack (HPL) | Pre-built binary MP_LINPACK INTEL - 2.3 |
STREAM | |
High Performance Conjugate Gradient (HPCG) | Pre-built binary from INTEL oneAPI 2.3 |
Ohio State University (OSU) |
In the present study, synthetic applications such as HPL, STREAM, and HPCG are done on a single node; since the OSU benchmark is a benchmark study on MPI operations, it requires a minimum of two nodes.
Synthetic application performance details
As shown in Table 2, four synthetic applications are tested on the test bed hardware (Table 1). They are HPL, STREAM, HPCG, and OSU. The details of performance of each application are given below:
High Performance Linpack (HPL)
HPL helps measure the floating-point computation efficiency of a system [1]. The details of the synthetic benchmarks can be found in the previous blog on Intel Ice Lake processors.
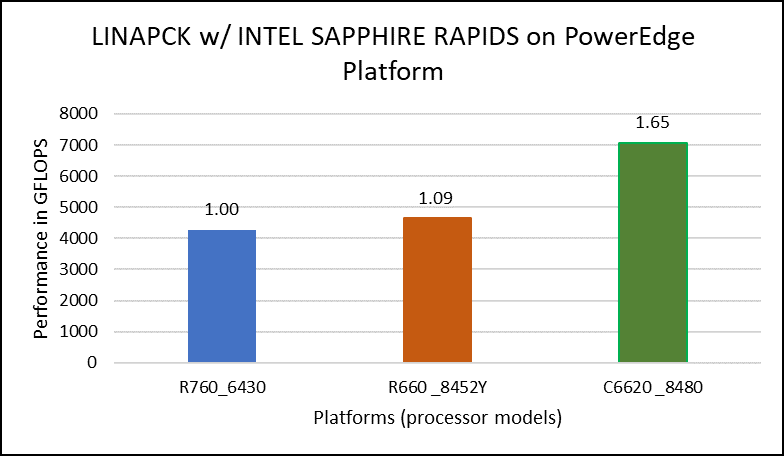
Figure 1. Performance values of HPL application for different processor models
The N and NB sizes used for the HPL benchmark are 348484 and 384, respectively, for the Intel Sapphire Rapids 6430, 8452Y processors, and 246144 and 384, respectively, for the 8480 processor. The difference in N sizes is due to the difference in available memory. Systems with Intel 6430 and 8452Y processors are equipped with 1024 GB of memory; the 8480 processor system has 512 GB. The performance numbers are captured with different BIOS settings, as discussed above, and the delta difference between each result is within 1-2%. The results with the HPC workload BIOS profile are shown in Figure 1. the 8452Y processor performs 1.09 times better than the Intel Sapphire Rapids 6430 processor and the 8480 processor performs 1.65 times better.
STREAM
The STREAM benchmark helps for measuring sustainable memory bandwidth of a processor. In general for STREAM benchmark, each array for STREAM must have at least four times the total size of all last-level caches utilized in the run or 1 million elements, whichever is larger. The STREAM array sizes used for the current study are 4×107 and 12×107 with full core utilization. The STREAM benchmark was also tested with 15 BIOS combinations, and the results depicted in Figure 2 are for the HPC workload profile bios test case. The STREAM TRIAD results are captured here in GB/sec. Results show improvement in performance compared to the Intel 3rd Generation Xeon Scalable processors, such as the 8380 and 6338. Also, if comparing 6430, 8452Y and 8480 processors, the STREAM results with 8452Y and 8480 Intel 4th Generation Xeon Scalable processors are, respectively, 1.12 and 1.24 times better than the Intel 6430 processor.
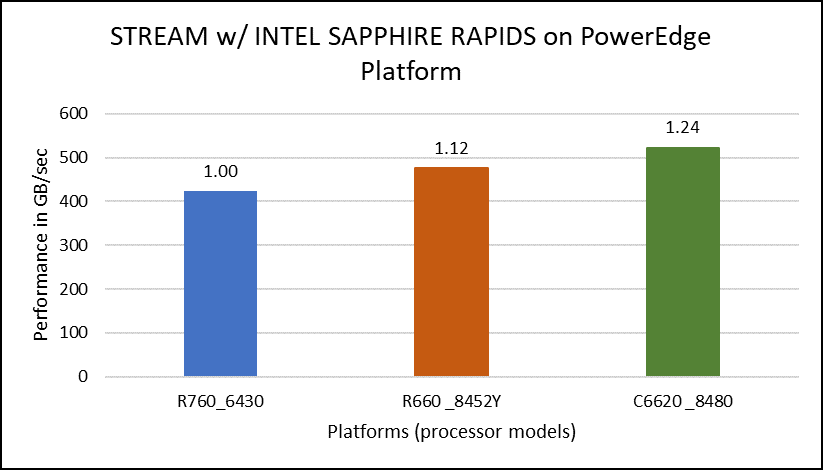
Figure 2. Performance values of STREAM application for different processor models
HPCG
The HPCG benchmark aims to simulate the data access patterns of applications such as sparse matrix calculations, assessing the impact of memory subsystem and internal connectivity constraints on the computing performance of High-Performance Computers, or supercomputers. The different problem sizes used in the study are 192, 256, 176, 168, and so on. Additionally, in this benchmark study, the variation in performance within different BIOS options was within 1–2 percent. Figure 3 shows the HPCG performance results for Intel Sapphire Rapids processors 6430, 8452Y and 8480. In comparison with the Intel 6430 processor, the 8452Y shows 1.02 times and the 8480 shows 1.12 times better performance.
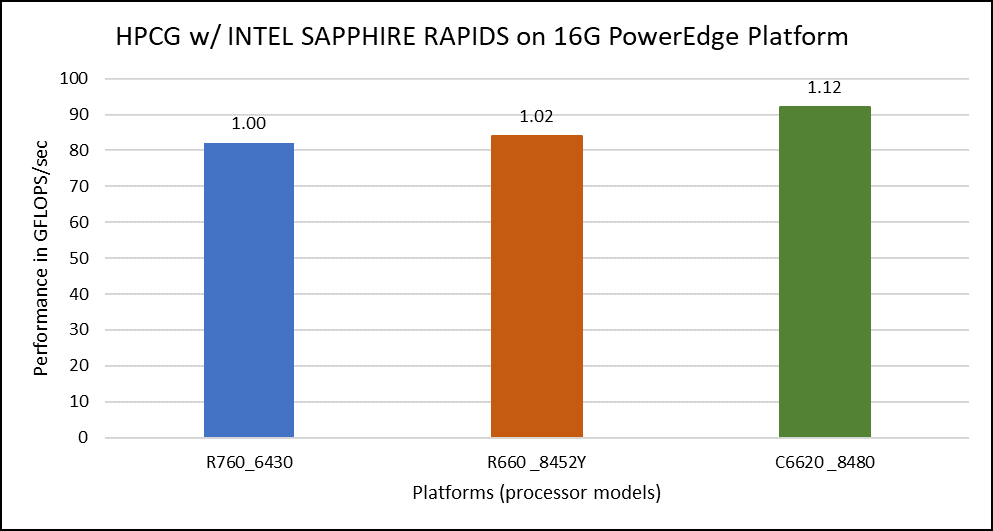
Figure 3. Performance values of HPCG application for different processor models
OSU Micro Benchmarks
OSU Micro Benchmarks are used for measuring the performance of MPI implementations, so we used two nodes connected to NDR200. OSU benchmark determines uni-directional and bi-directional bandwidth and message rate and latency between the nodes. The OSU benchmark was run on all three Intel processors (6430, 8452Y, and 8480) with single core per node; however, we have shown one of the system/processors (Intel 8480 processor) results in the blog starting from Figures 4-7.
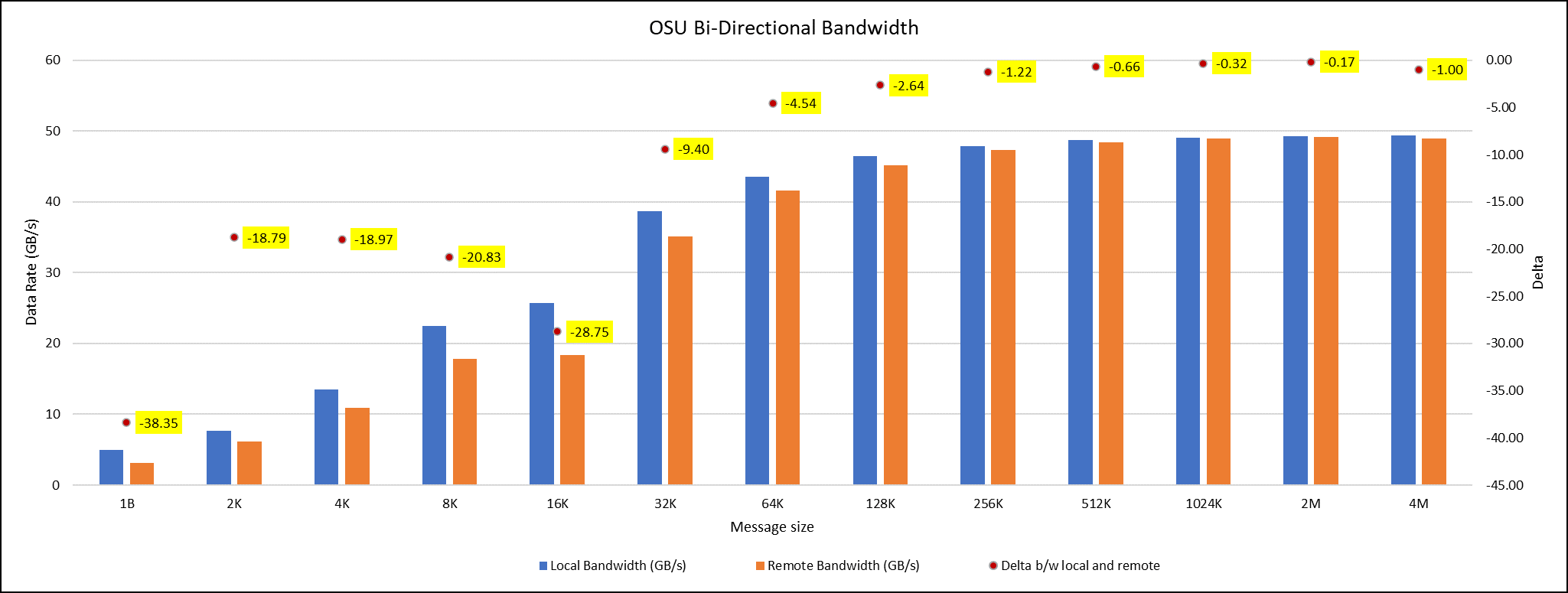
Figure 4. OSU Bi-Directional bandwidth chart for C6620_8480 intel processor
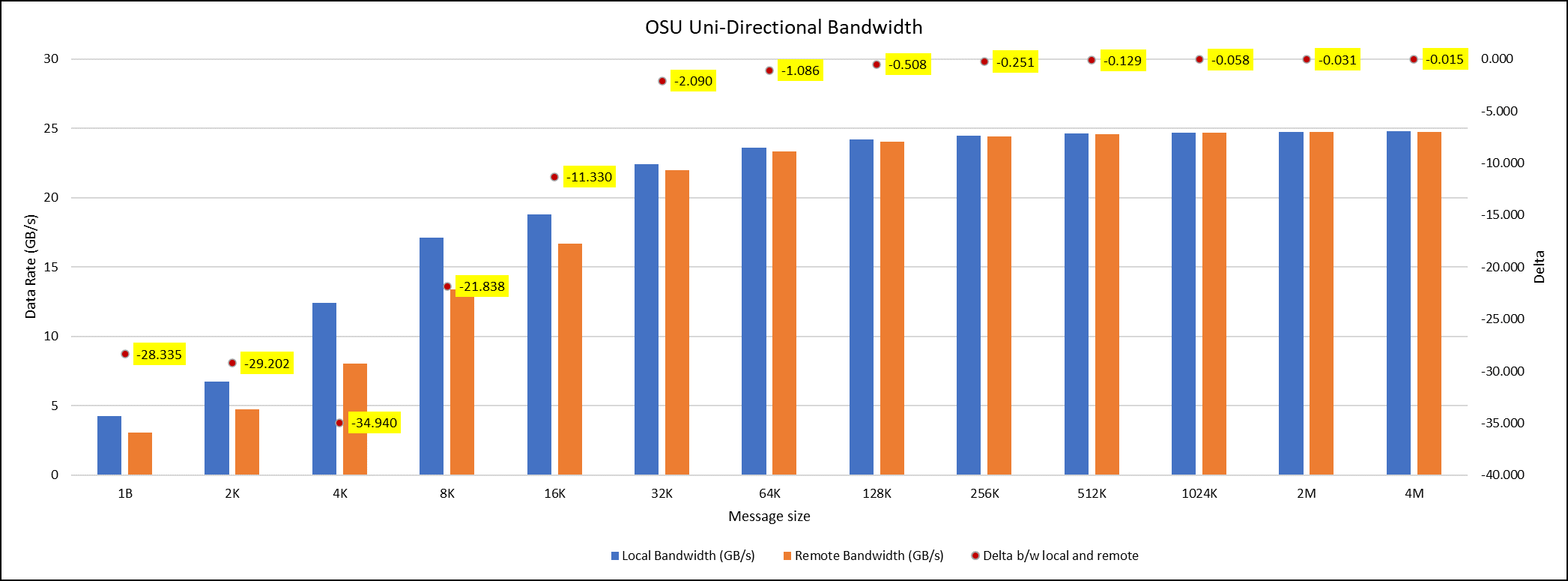
Figure 5. OSU Uni-Directional bandwidth chart for C6620_8480 intel processor
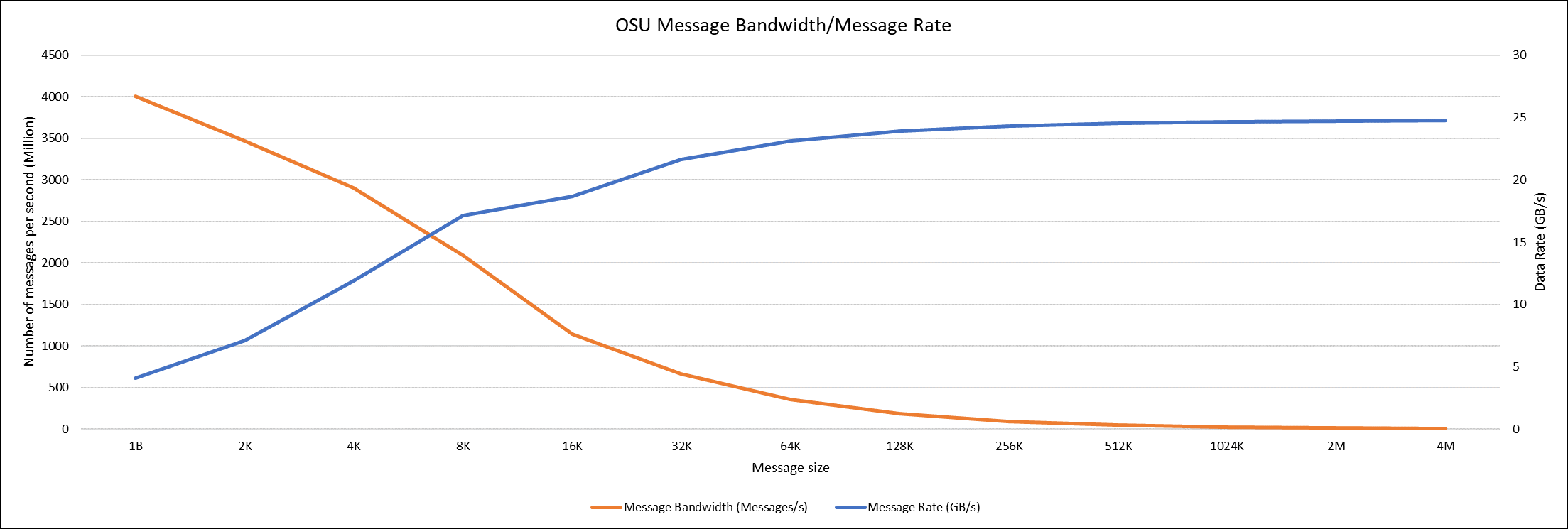
Figure 6. OSU Message bandwidth/Message rate chart for C6620_8480 intel processor
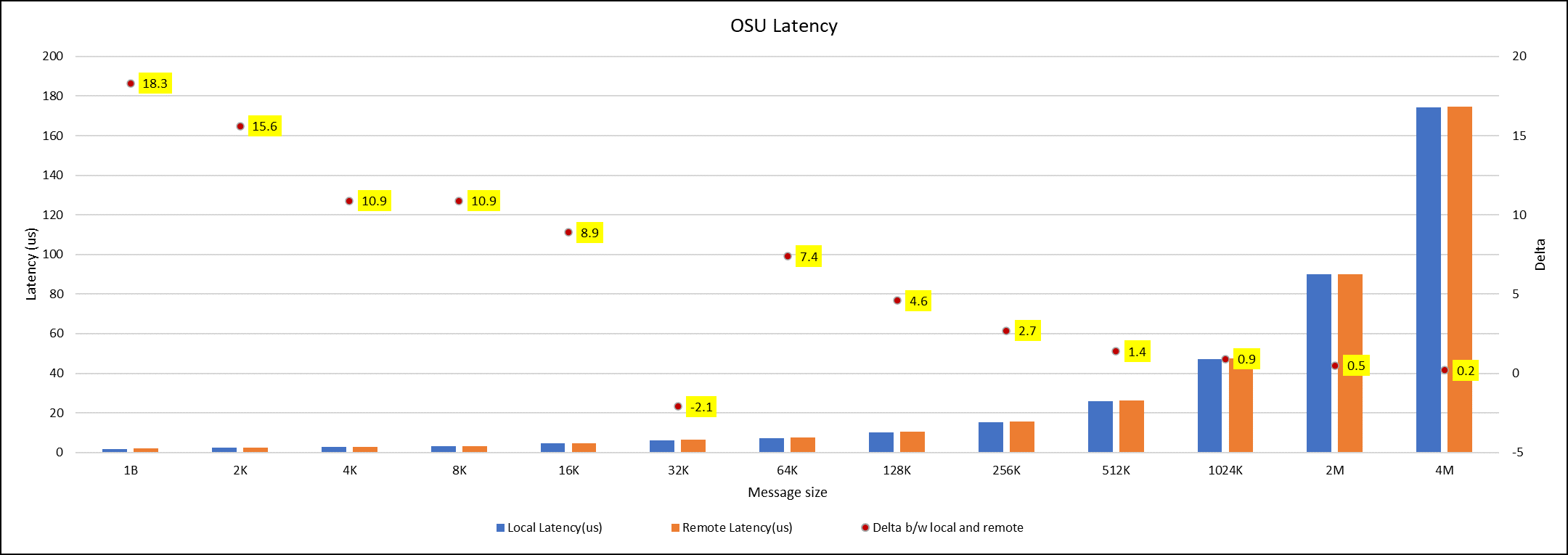
Figure 7. OSU Latency chart for C6620_8480 intel processor
All fifteen BIOS combinations were tested; the OSU benchmark also shows similar performance with a difference within a 1-2% delta.
Conclusion
The performance comparison between various Intel Sapphire Rapids processors (6430, 8452Y and 8480) is done with the help of synthetic benchmark applications such as HPL, STREAM, HPCG and OSU. Nearly 15 BIOS configurations are set on the system, and performance values with different benchmarks were captured to identify the best BIOS configuration to set. From the results, it was found that the difference in performance with any benchmarks for all the BIOS configurations applied is below 3 percent delta.
Therefore, the HPC workload profile provides better benchmark results with all the Intel Sapphire Rapids processors. Among the three Intel processors compared, the 8480 had the highest application performance value, while the 8452Y is in second place. The maximum difference in performance between processors was found for the HPL benchmark, and it was the 8480 Intel Sapphire Rapids processor, which offers 1.65 times better results than the Intel 6430 processor.
Watch out for future application benchmark results on this blog! Visit our page for previous blogs.


