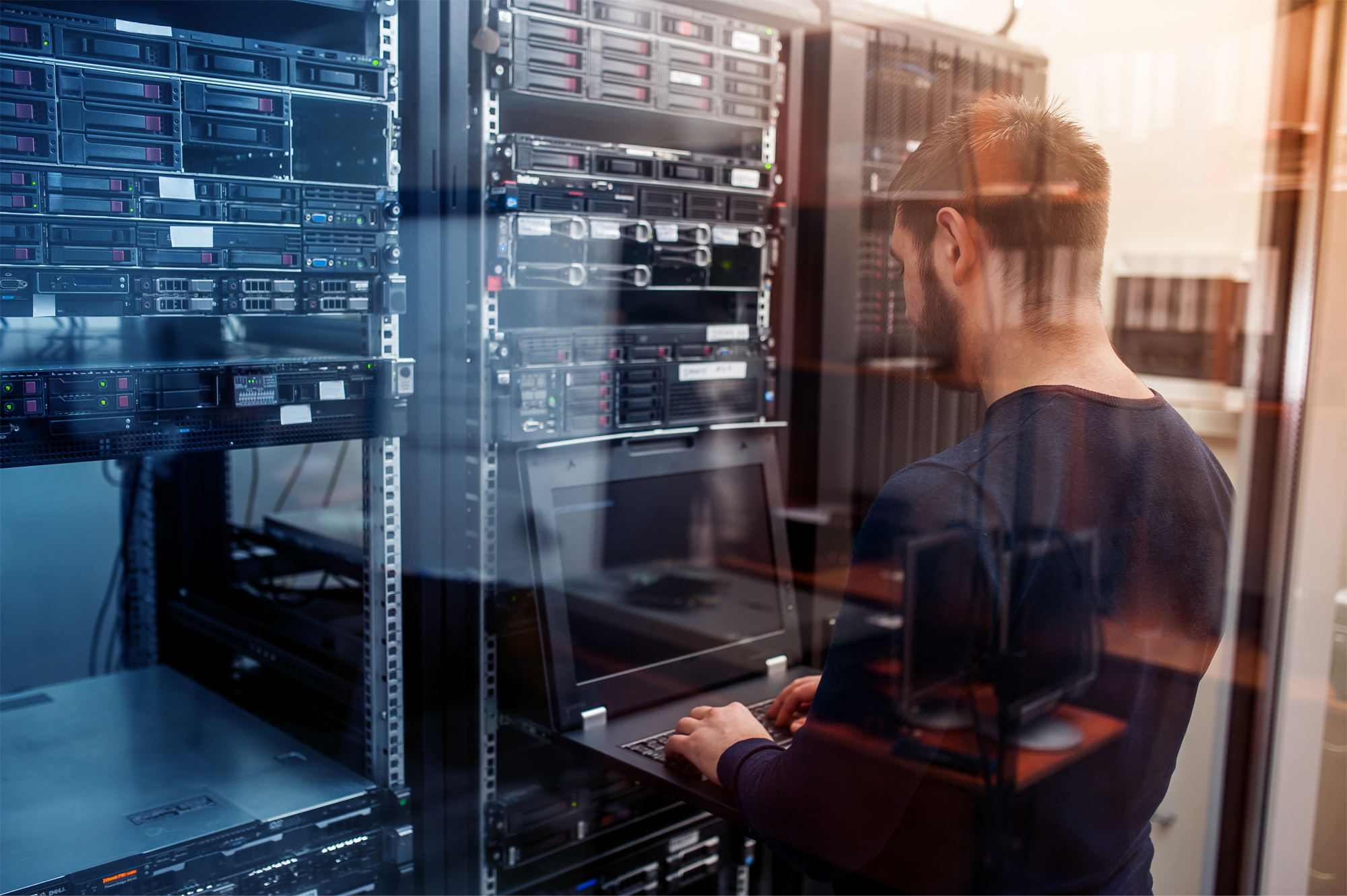


Tuxedo Pipeline Performance on Dell EMC PowerEdge R6525
Tue, 27 Apr 2021 03:48:30 -0000
|Read Time: 0 minutes
Overview
Gene expression analysis is as important as identifying Single Nucleotide Polymorphism (SNP), insertion/deletion (indel), or chromosomal restructuring. Ultimately, all physiological and biochemical events depend on the final gene expression products, proteins. Although most mammals have an additional controlling layer before protein expression, knowing how many transcripts exist in a system helps to characterize the biochemical status of a cell. Ideally, technology would enable us to quantify all proteins in a cell, which would significantly advance the progress of Life Science. However, we are far from achieving this.
This blog provides the test results of one popular RNA-Seq data analysis pipeline known as the Tuxedo pipeline (1). The Tuxedo pipeline suite offers a set of tools for analyzing a variety of RNA-Seq data, including short-read mapping, identification of splice junctions, transcript, and isoform detection, differential expression, visualizations, and quality control metrics. The tested workflow is a differentially expressed gene (DEG) analysis, and the detailed steps in the pipeline are shown in Figure 1.
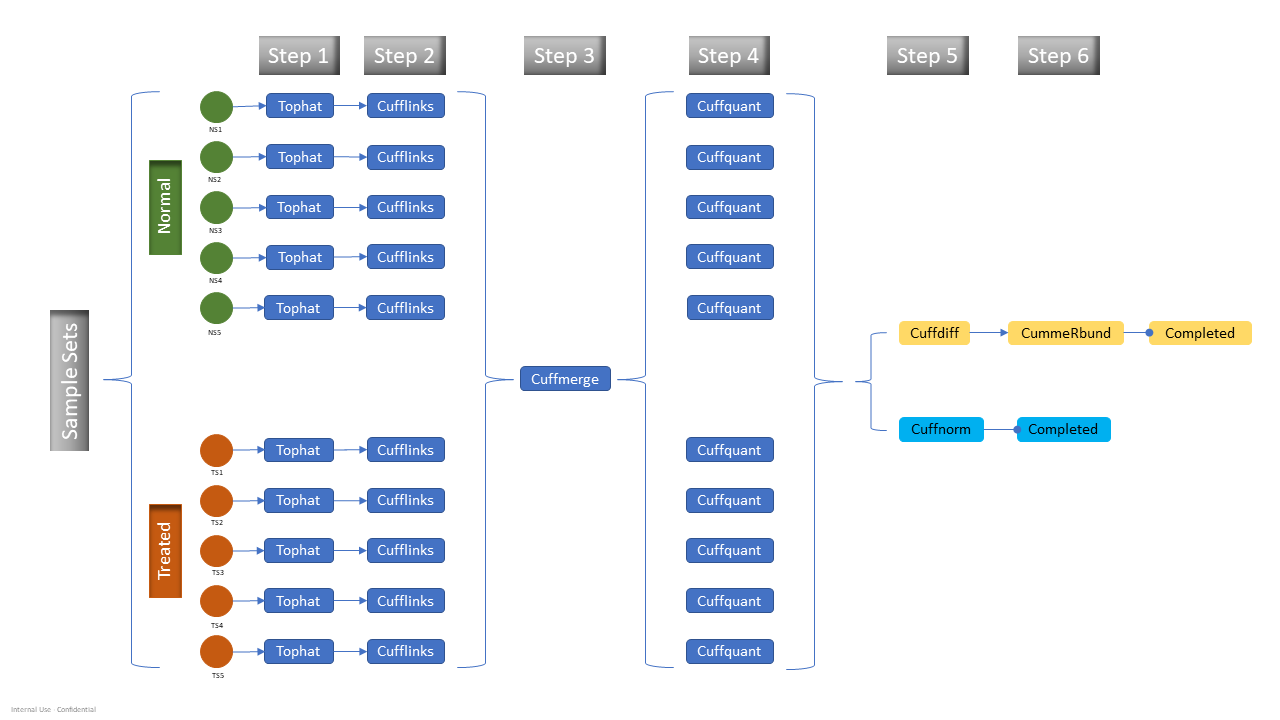
Figure 1. Updated tuxedo pipeline with cuffquant step
A single node study with AMD EPYC 7002 series (Rome) and AMD EPYC 7003 series (Milan) on Dell EMC PowerEdge R6525 server was done. The configurations of the test system are summarized in Table 1.
Table 1. Tested compute node configuration
| Dell EMC PowerEdge R6525 | |
|---|---|
| CPU | Tested AMD Milan: 2x 7763 (Milan), 64 Cores, 2.45 GHz – 3.5 GHz Base-Boost, TDP 280 W, 256 MB L3 Cache 2x 7713 (Milan), 64 Cores, 2.0 GHz – 3.7 GHz Base-Boost, TDP 225 W, 256 MB L3 Cache 7543 (Milan), 32 Cores, 2.8 GHz – 3.7 GHz Base-Boost, TDP 225 W, 256 MB L3 Cache
Tested AMD Rome: 7702 (Rome), 64 Cores, 2.0 GHz – 3.35 GHz Base-Boost, TDP 200 W, 256 MB L3 Cache |
| RAM | DDR4 256 GB (16 Gb x 16) 3200 MT/s |
| Operating system | RHEL 8.3 (4.18.0-240.el8.x86_64) |
| Interconnect | Mellanox InfiniBand HDR100 |
| Filesystem | Dell EMC Ready Solutions for HPC BeeGFS High Capacity Storage |
| BIOS system profile | Performance optimized |
| Logical processor | Disabled |
| Virtualization technology | Disabled |
| tophat | 2.1.1 |
| bowtie2 | 2.2.5 |
| R | 3.6 |
| bioconductor cummerbund | 2.26.0 |
A performance study of the RNA-Seq pipeline is not trivial because the nature of the workflow requires non-identical input files yet similar input files in size. Hence, 185 RNA-Seq paired-end read data are collected from a public data repository. All the read datafiles contain around 25 Million Fragments (MF) and have similar read lengths. The samples for a test are randomly selected from the pool of the 185 paired-end read files. Although these test data will not have any biological meaning, certainly these data with the high level of noise will put the tests in the worst-case scenario.
Performance Evaluation
Throughput test - Single pipeline with more than two samples, biological, and technical duplicates
Typical RNA-Seq studies consist of multiple samples, sometimes 100s of different samples, normal versus disease, or untreated versus treated samples. These samples tend to have a high level of noise due to biological reasons; hence, the analysis requires vigorous data preprocessing procedure.
A number of various samples were processed, with different RNA-Seq data selected from 185 paired-end reads dataset, to see how much data a single node can process. Typically, when the number of samples increases, the runtime of the Tuxedo pipeline increases. However, as shown in the figure below, the runtimes with two sample tests using 7713, are higher than the runtimes from four samples. The standard error from five repeated runs does not overlap with four and eight sample results. The interference of other users may cause this large variance. The current testing environment, especially a shared file system designed for large capacity, is not ideal for a Next Generation Sequencing (NGS) data analysis benchmark.
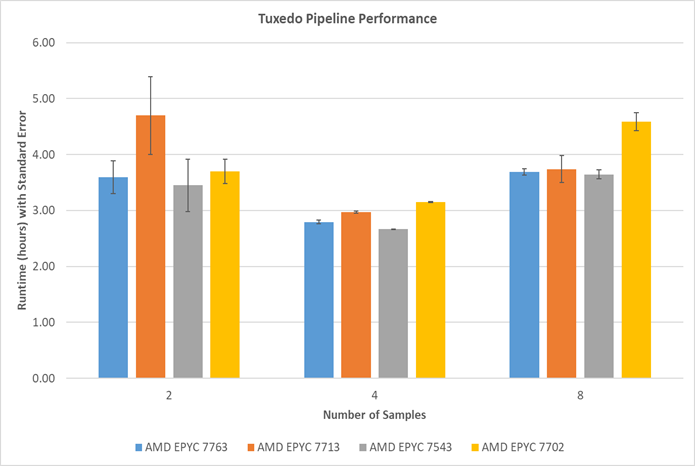
Figure 2. Runtime comparisons among various AMD EPYC 7003 Series processors: Standard error is estimated from an estimated standard deviation based on a sample (STDDEV.S function in Excel)
The eight sample test results show that AMD Milan processors perform better than one of the Rome processors (7702) in a higher workload.
Conclusion
Many tests are still required to obtain a better insight from the AMD Milan processors for the NGS data analysis area. Unfortunately, the tests could not exceed eight samples due to storage limitations. However, there seems to be plenty of room for a higher throughput that processes more than eight samples together. AMD Milan 7763 performed 20% better than AMD Rome 7702. AMD Milan 7713 performed 18% better in eight sample tests for the Tuxedo pipeline as described in Figure 2.
Related Blog Posts

HPC Application Performance on Dell PowerEdge R7525 Servers with the AMD Instinct™ MI210 GPU
Mon, 12 Sep 2022 12:11:52 -0000
|Read Time: 0 minutes
PowerEdge support and performance
The PowerEdge R7525 server can support three AMD Instinct™ MI210 GPUs; it is ideal for HPC Workloads. Furthermore, using the PowerEdge R7525 server to power AMD Instinct MI210 GPUs (built with the 2nd Gen AMD CDNA™ architecture) offers improvements on FP64 operations along with the robust capabilities of the AMD ROCm™ 5 open software ecosystem. Overall, the PowerEdge R7525 server with the AMD Instinct MI210 GPU delivers expectational double precision performance and leading total cost of ownership.

Figure 1: Front view of the PowerEdge R7525 server
We performed and observed multiple benchmarks with AMD Instinct MI210 GPUs populated in a PowerEdge R7525 server. This blog shows the performance of LINPACK and the OpenMM customizable molecular simulation libraries with the AMD Instinct MI210 GPU and compares the performance characteristics to the previous generation AMD Instinct MI100 GPU.
The following table provides the configuration details of the PowerEdge R7525 system under test (SUT):
Table 1. SUT hardware and software configurations
Component | Description |
Processor | AMD EPYC 7713 64-Core Processor |
Memory | 512 GB |
Local disk | 1.8T SSD |
Operating system | Ubuntu 20.04.3 LTS |
GPU | 3xMI210/MI100 |
Driver version | 5.13.20.22.10 |
ROCm version | ROCm-5.1.3 |
Processor Settings > Logical Processors | Disabled |
System profiles | Performance |
NUMA node per socket | 4 |
HPL | rochpl_rocm-5.1-60_ubuntu-20.04 |
OpenMM | 7.7.0_49 |
The following table contains the specifications of AMD Instinct MI210 and MI100 GPUs:
Table 2: AMD Instinct MI100 and MI210 PCIe GPU specifications
GPU architecture | AMD Instinct MI210 | AMD Instinct MI100 |
Peak Engine Clock (MHz) | 1700 | 1502 |
Stream processors | 6656 | 7680 |
Peak FP64 (TFlops) | 22.63 | 11.5 |
Peak FP64 Tensor DGEMM (TFlops) | 45.25 | 11.5 |
Peak FP32 (TFlops) | 22.63 | 23.1 |
Peak FP32 Tensor SGEMM (TFlops) | 45.25 | 46.1 |
Memory size (GB) | 64 | 32 |
Memory Type | HBM2e | HBM2 |
Peak Memory Bandwidth (GB/s) | 1638 | 1228 |
Memory ECC support | Yes | Yes |
TDP (Watt) | 300 | 300 |
High-Performance LINPACK (HPL)
HPL measures the floating-point computing power of a system by solving a uniformly random system of linear equations in double precision (FP64) arithmetic, as shown in the following figure. The HPL binary used to collect results was compiled with ROCm 5.1.3.
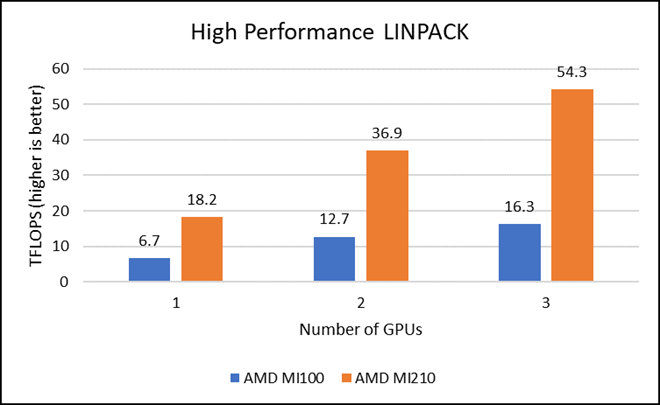
Figure 2: LINPACK performance with AMD Instinct MI100 and MI210 GPUs
The following figure shows the power consumption during a single HPL run:
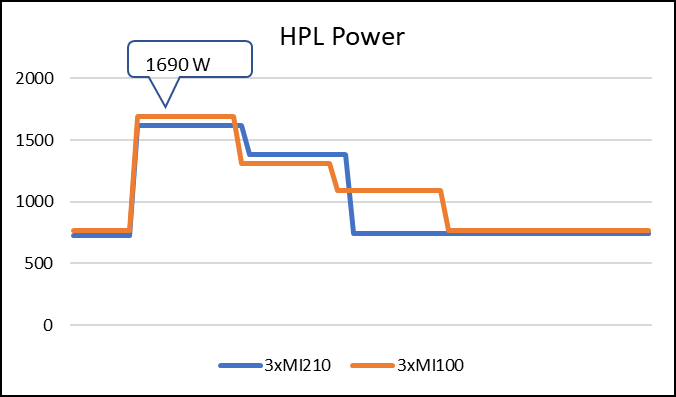
Figure 3: LINPACK power consumption with AMD Instinct MI100 and MI210 GPUs
We observed a significant improvement in the AMD Instinct MI210 HPL performance over the AMD Instinct MI100 GPU. The numbers on a single GPU test of MI210 are 18.2 TFLOPS which is approximately 2.7 times higher than MI100 number (6.75 TFLOPS). This improvement is due to the AMD CDNA2 architecture on the AMD Instinct MI210 GPU, which has been optimized for FP64 matrix and vector workloads. Also, the MI210 GPU has larger memory, so the problem size (N) used here is large in comparison to the AMD Instinct MI100 GPU.
As shown in Figure 2, the AMD Instinct MI210 has shown almost linear scalability in the HPL values on single node multi-GPU runs. The AMD Instinct MI210 GPU reports better scalability compared to its last generation AMD Instinct MI100 GPUs. Both GPUs have the same TDP, with the AMD Instinct MI210 GPU delivering three times better performance. The performance per watt value of a PowerEdge R7525 system is three times more. Figure 3 shows the power consumption characteristics in one HPL run cycle.
OpenMM
OpenMM is a high-performance toolkit for molecular simulation. It can be used as a library or as an application. It includes extensive language bindings for Python, C, C++, and even Fortran. The code is open source and actively maintained on GitHub and licensed under MIT and LGPL.
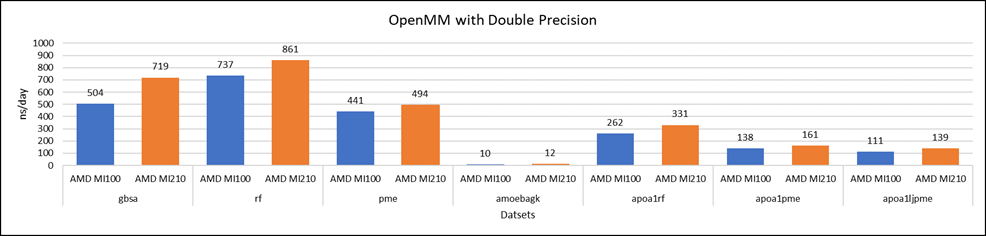
Figure 4: OpenMM double-precision performance with AMD Instinct MI100 and MI210 GPUs
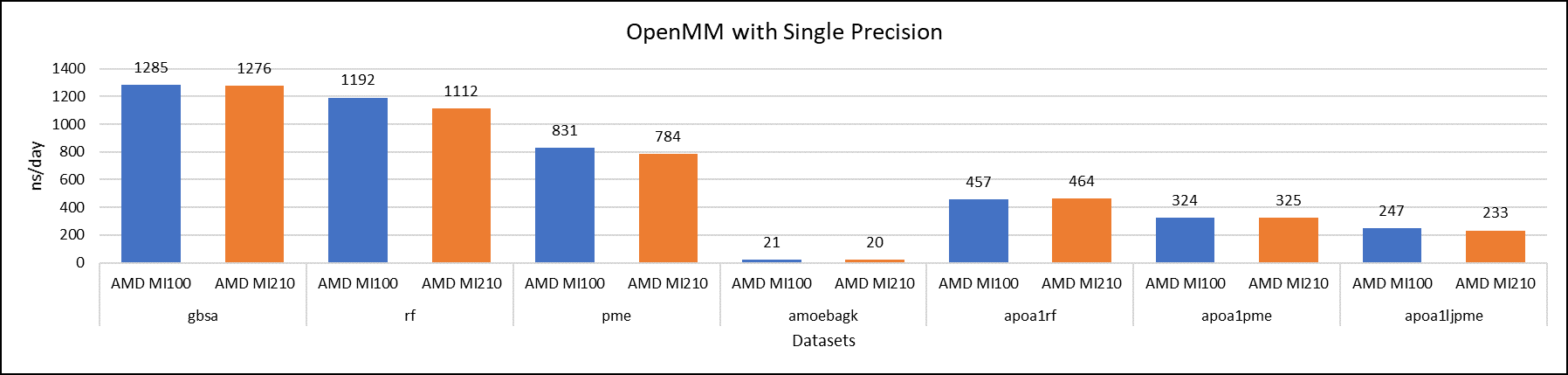
Figure 5: OpenMM single-precision performance with AMD Instinct MI100 and MI210 GPUs
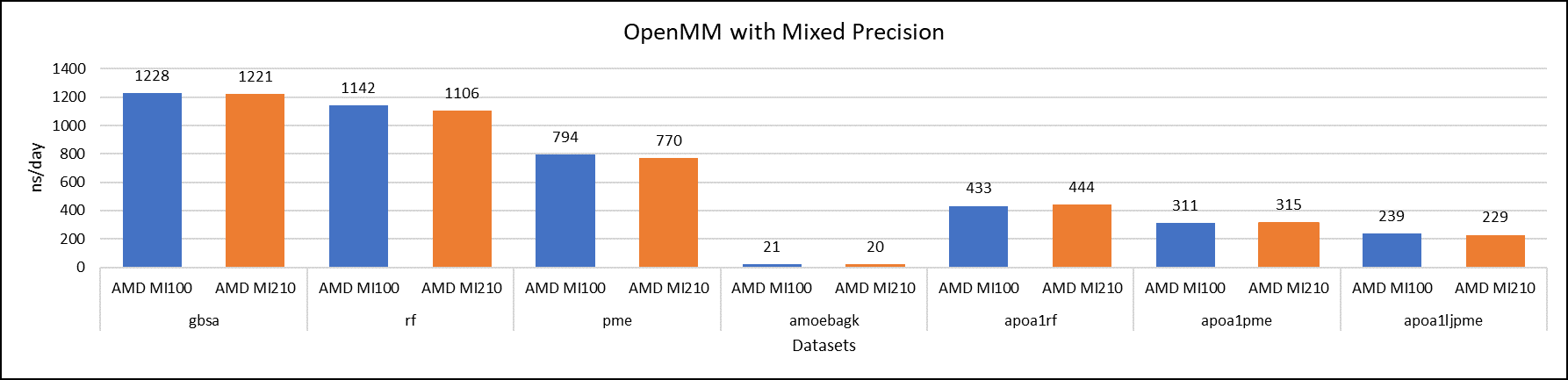
Figure 6: OpenMM mixed-precision performance with AMD Instinct MI100 and MI210 GPUs
We tested OpenMM with seven datasets to validate double, single, and mixed precision. We observed exceptional double precision performance with OpenMM on the AMD Instinct MI210 GPU compared to the AMD Instinct MI100 GPU. This improvement is due to the AMD CDNA2 architecture on the AMD Instinct MI210 GPU, which has been optimized for FP64 matrix and vector workloads.
Conclusion
The AMD Instinct MI210 GPU shows an impressive performance improvement in FP64 workloads. These workloads benefit as AMD has doubled the width of their ALUs to a full 64-bits wide. This change allows the FP64 operations to now run at full speed in the new 2nd Gen AMD CDNA architecture. The applications and workloads that are designed to run on FP64 operations are expected to take full advantage of the hardware.

HPC Application Performance on Dell PowerEdge R750xa Servers with the AMD Instinct TM MI210 Accelerator
Fri, 12 Aug 2022 16:47:40 -0000
|Read Time: 0 minutes
Overview
The Dell PowerEdge R750xa server, powered by 3rd Generation Intel Xeon Scalable processors, is a 2U rack server that supports dual CPUs, with up to 32 DDR4 DIMMs at 3200 MT/s in eight channels per CPU. The PowerEdge R750xa server is designed to support up to four PCI Gen 4 accelerator cards and up to eight SAS/SATA SSD or NVMe drives.

Figure 1: Front view of the PowerEdge R750xa server
The AMD Instinct™ MI210 PCIe accelerator is the latest GPU from AMD that is designed for a broad set of HPC and AI applications. It provides the following key features and technologies:
- Built with the 2nd Gen AMD CDNA architecture with new Matrix Cores delivering improvements on FP64 operations and enabling a broad range of mixed-precision capabilities
- 64 GB high-speed HBM2e memory bandwidth supporting highly data-intensive workloads
- 3rd Gen AMD Infinity Fabric™ technology bringing advanced platform connectivity and scalability enabling fully connected dual P2P GPU hives through AMD Infinity Fabric™ links
- Combined with the AMD ROCm™ 5 open software platform allowing researchers to tap the power of the AMD Instinct™ accelerator with optimized compilers, libraries, and runtime support
This blog provides the performance characteristics of a single PowerEdge R750xa server with the AMD Instinct MI210 accelerator. It compares the performance numbers of microbenchmarks (GEMM of FP64 and FP32 and bandwidth test), HPL, and LAMMPS for both the AMD Instinct MI210 accelerator and the previous generation AMD Instinct MI100 accelerator.
The following table provides configuration details for the PowerEdge R750xa system under test (SUT):
Table 1: SUT hardware and software configurations
Component | Description |
Processor | Dual Intel Xeon Gold 6338 |
Memory | 512 GB - 16 x 32 GiB@3200 MHz |
Local disk | 3.84 TB SATA-6GB SSD |
Operating system | Rocky Linux release 8.4 (Green Obsidian) |
GPU model | 4 x AMD MI210 (PCIe-64G) or 3 x AMD MI100 (PCIe-32G) |
GPU driver version | 5.13.20.5.1 |
ROCm version | 5.1.3 |
Processor Settings > Logical Processors | Disabled |
System profiles | Performance |
5.1.3 | |
5.1.3 | |
HPL | Compiled with ROCm v5.1.3 |
LAMMPS (KOKKOS) | Version: LAMMPS patch_4May2022 |
The following table provides the specifications of the AMD Instinct MI210 and MI100 GPUs:
Table 2: AMD Instinct MI100 and MI210 PCIe GPU specifications
GPU architecture | AMD Instinct MI210 | AMD Instinct MI100 |
Peak Engine Clock (MHz) | 1700 | 1502 |
Stream processors | 6656 | 7680 |
Peak FP64 (TFlops) | 22.63 | 11.5 |
Peak FP64 Tensor DGEMM (TFlops) | 45.25 | 11.5 |
Peak FP32 (TFlops) | 22.63 | 23.1 |
Peak FP32 Tensor SGEMM (TFlops) | 45.25 | 46.1 |
Memory size (GB) | 64 | 32 |
Memory Type | HBM2e | HBM2 |
Peak Memory Bandwidth (GB/s) | 1638 | 1228 |
Memory ECC support | Yes | Yes |
TDP (Watt) | 300 | 300 |
GEMM microbenchmarks
Generic Matrix-Matrix Multiplication (GEMM) is a multithreaded dense matrix multiplication benchmark that is used to measure the performance of a single GPU. The unique O(n3) computational complexity compared to the O(n2) memory requirement of GEMM makes it an ideal benchmark to measure GPU acceleration with high efficiency because achieving high efficiency depends on minimizing the redundant memory access.
For this test, we complied the rocblas-bench binary from https://github.com/ROCmSoftwarePlatform/rocBLAS to collect DGEMM (double-precision) and SGEMM (single-precision) performance numbers.
These results only reflect the performance of matrix multiplication, and results are measured in the form of peak TFLOPS that the accelerator can deliver. These numbers can be used to compare the peak compute performance capabilities of different accelerators. However, they might not represent real-world application performance.
Figure 2 presents the performance results measured for DGEMM and SGEMM on a single GPU:
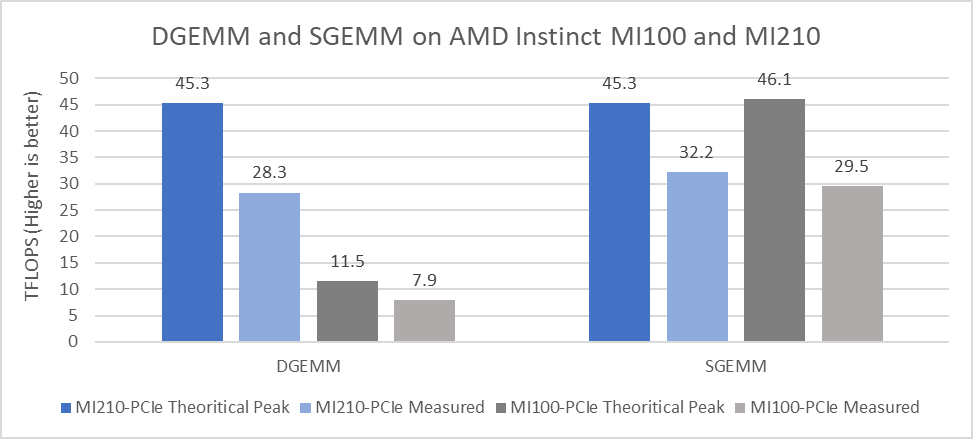
Figure 2: DGEMM and SGEMM numbers obtained on AMD Instinct MI210 and MI100 GPUs with the PowerEdge R750xa server
From the results we observed:
- The CDNA 2 architecture from AMD, which includes second-generation Matrix Cores and faster memory, provides significant improvement in the theoretical peak FP64 Tensor DGEMM value (45.3 TFLOPS). This result is 3.94 times better than the previous generation AMD Instinct MI100 GPU peak of 11.5 TFLOPS. The measured DGEMM value on the AMD Instinct MI250 GPU is 28.3 TFlops, which is 3.58 times better compared to the measured value of 7.9 TFlops on the AMD Instinct MI100 GPU.
- For FP32 Tensor operations in the SGEMM single-precision GEMM benchmark, the theoretical peak performance of the AMD Instinct MI210 GPU is 45.23 TFLOPS, and the measured performance value is 32.2 TFLOPS. An improvement of approximately nine percent was observed in the measured value of SGEMM compared to the AMD Instinct MI100 GPU.
GPU-to-GPU bandwidth test
This test captures the performance characteristics of buffer copying and kernel read/write operations. We collected results by using TransferBench, compiling the binary by following the procedure provided at https://github.com/ROCmSoftwarePlatform/rccl/tree/develop/tools/TransferBench. On the PowerEdge R750xa server, both the AMD Instinct MI100 and MI210 GPUs have the same GPU-to-GPU throughput, as shown in the following figure:
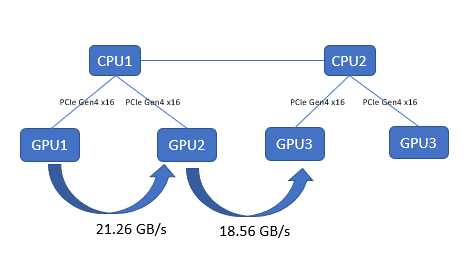
Figure 3: GPU-to-GPU bandwidth test with TransferBench on the PowerEdge R750xa server with AMD Instinct MI210 GPUs
High-Performance Linpack (HPL) Benchmark
HPL measures a system’s floating point computing power by solving a random system of linear equations in double precision (FP64) arithmetic. The peak FLOPS (Rpeak) is the highest number of floating-point operations that a computer can perform per second in theory.
It can be calculated using the following formula:
clock speed of the GPU × number of GPU cores × number of floating-point operations that the GPU performs per cycle
Measured performance is referred to as Rmax. The ratio of Rmax to Rpeak demonstrates the HPL efficiency, which is how close the measured performance is to the theoretical peak. Several factors influence efficiency including GPU core clock speed boost and the efficiency of the software libraries.
The results shown in the following figure are the Rmax values, which are measured HPL numbers on AMD Instinct MI210 and AMD MI100 GPUs. The HPL binary used to collect the result was compiled with ROCm 5.1.3.
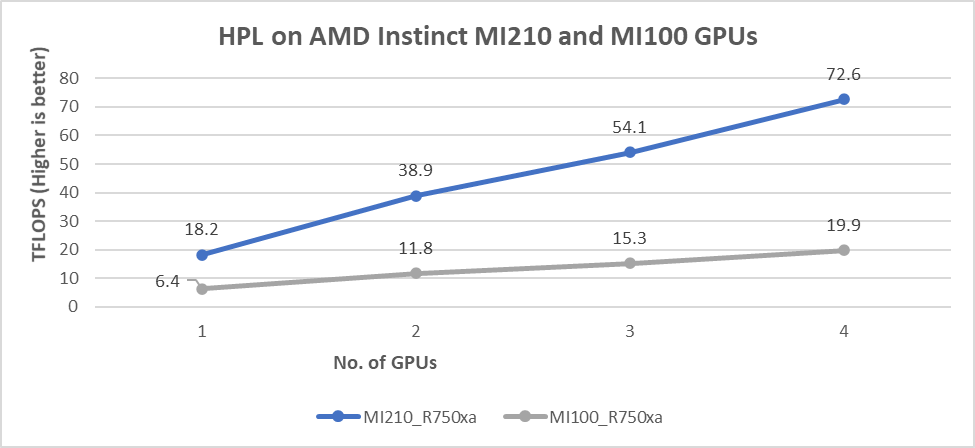
Figure 4: HPL performance on AMD Instinct MI210 and MI100 GPUs powered with R750xa servers
The following figure shows the power consumption during a single HPL test :
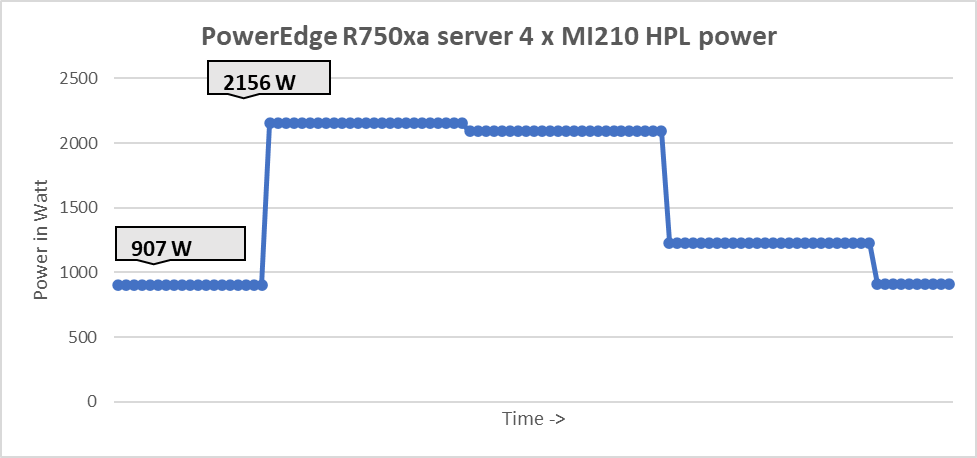

Figure 5: System power use during one HPL test across four GPUs
Our observations include:
- We observed a significant improvement in the HPL performance with the AMD Instinct MI210 GPU over the AMD Instinct MI100 GPU. The performance on a single test of the AMD Instinct MI210 GPU is 18.2 TFLOPS, which is over 2.8 times higher than the AMD Instinct MI100 number of 6.4 TFLOPS. This improvement is a result of the AMD CDNA2 architecture on the AMD Instinct MI210 GPU, which has been optimized for FP64 matrix and vector workloads.
- As shown in Figure 4, the AMD Instinct MI210 GPU provides almost linear scalability in the HPL values on single node multi-GPU runs. The AMD Instinct MI210 GPU shows better scalability compared to the previous generation AMD Instinct MI100 GPUs.
- Both AMD Instinct MI100 and MI210 GPUs have the same TDP of 300 W, with the AMD Instinct MI210 GPU delivering a 3.6 times better performance. The performance per watt value from a PowerEdge R750xa server is 3.6 times more.
LAMMPS Benchmark
LAMMPS is a molecular dynamics simulation code that is a GPU bandwidth-bounded application. We used the KOKKOS acceleration library implementation of LAMMPS to measure the performance of AMD Instinct MI210 GPUs.
The following figure compares the LAMMPS performance of the AMD Instinct MI210 and MI100 GPU with four different datasets:
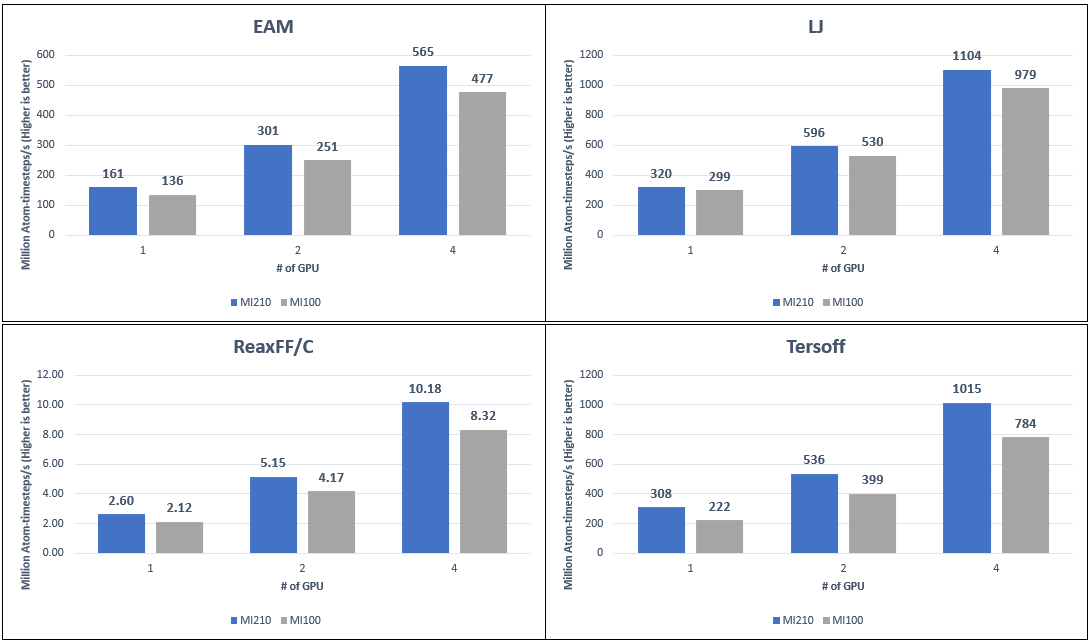
Figure 6: LAMMPS performance numbers on AMD Instinct MI210 and MI100 GPUs on PowerEdge R750xa servers with different datasets
Our observations include:
- We measure to an average 21 percent performance improvement on the AMD Instinct MI210 GPU compared to the AMD Instinct MI100 GPU with the PowerEdge R750xa server. Because MI100 and MI210 GPUs have different sizes of onboard GPU memory, the problem sizes of each LAMMPS dataset were adjusted to represent the best performance from each GPU.
- Datasets such as Tersoff, ReaxFF/C, and EAM on the AMD Instinct MI210 GPU show a 30 percent, 22 percent, and 18 percent improvement. This result is primarily because the AMD Instinct MI210 GPU comes with faster and larger memory HBM2e (64 GB) compared to the AMD Instinct MI100 GPU, which comes with HBM2 (32 GB) memory. For the LJ datasets, the improvement is less, but is still observed at 12 percent. This result is because single-precision calculations are used and the FP32 peak performance for the AMD Instinct MI210 and MI100 GPUs are at the same level.
Conclusion
The AMD Instinct MI210 GPU shows impressive performance improvement in FP64 workloads. These workloads benefit as AMD has doubled the width of their ALUs to a full 64 bits wide allowing FP64 operations to now run at full speed in the new CDNA 2 architecture. Applications and workloads that can take advantage of FP64 operations are expected to make the most of the aspect of the AMD Instinct MI210 GPU. The faster bandwidth of the HBM2e memory of the AMD Instinct MI210 GPU provides advantages for GPU memory-bounded applications.
The PowerEdge R750xa server with AMD Instinct MI210 GPUs is a powerful compute engine, which is well suited for HPC users who need accelerated compute solutions.
Next steps
In future work, we plan to describe benchmark results on additional HPC and deep learning applications, compare the AMD Infinity FabricTM Link(xGMI) bridges, and show AMD Instinct MI210 performance numbers on other Dell PowerEdge servers, such as the PowerEdge R7525 server.


