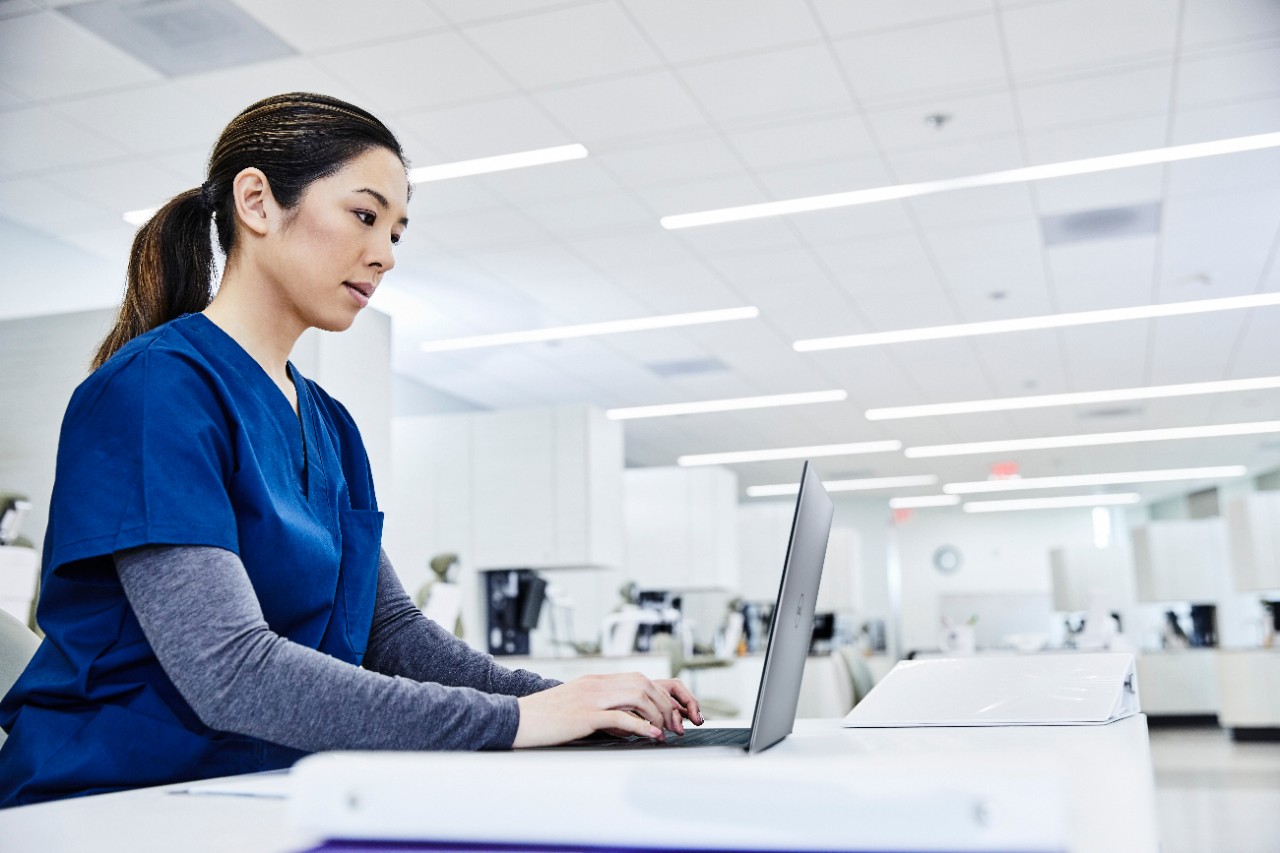
MD Simulation of GROMACS with AMD EPYC 7003 Series Processors on Dell EMC PowerEdge Servers
Thu, 19 Aug 2021 20:06:53 -0000
|Read Time: 0 minutes
AMD has recently announced and launched its third generation 7003 series EPYC processors family (code named Milan). These processors build upon the proceeding generation 7002 series (Rome) processors and improve L3 cache architecture along with an increased memory bandwidth for workloads such as High Performance Computing (HPC).
The Dell EMC HPC and AI Innovation Lab has been evaluating these new processors with Dell EMC’s latest 15G PowerEdge servers and will report our initial findings for the molecular dynamics (MD) application GROMACs in this blog.
Given the enormous health impact of the ongoing COVID-19 pandemic, researchers and scientists are working closely with the HPC and AI Innovation Lab to obtain the appropriate computing resources to improve the performance of molecular dynamics simulations. Of these resources, GROMACS is an extensively used application for MD simulations. It has been evaluated with the standard datasets by combining the latest AMD EPYC Milan processor (based on Zen 3 cores) with Dell EMC PowerEdge servers to get most out of the MD simulations.
In a previous blog, Molecular Dynamic Simulation with GROMACS on AMD EPYC- ROME, we published benchmark data for a GROMACS application study on a single node and multinode with AMD EPYC ROME based Dell EMC servers.
The results featured in this blog come from the test bed described in the following table. We performed a single-node and multi-node application study on Milan processors, using the latest AMD stack shown in Table 1, with GROMACS 2020.4 to understand the performance improvement over the older generation processor (Rome).
Table 1: Testbed hardware and software details
Server | Dell EMC PowerEdge 2-socket servers (with AMD Milan processors) | Dell EMC PowerEdge 2-socket servers (with AMD Rome processors) |
Processor Cores/socket Frequency (Base-Boost ) Default TDP Processor bus speed | 7763 (Milan) 64 2.45 GHz – 3.5 GHz 280 W 256 MB 16 GT/s | 7H12 (Rome) 64 2.6 GHz – 3.3 GHz 280 W 256 MB 16 GT/s |
Processor Cores/socket Frequency Default TDP Processor bus speed | 7713 (Milan) 64 2.0 GHz – 3.675 GHz 225 W 256 MB 16 GT/s | 7702 (Rome) 64 2.0 GHz – 3.35 GHz 200 W 256 MB 16 GT/s |
Processor Cores/socket Frequency Default TDP Processor bus speed | 7543 (Milan) 32 2.8 GHz – 3.7 GHz 225 W 256 MB 16 GT/s | 7542 (Rome) 32 2.9 GHz – 3.4 GHz 225 W 128 MB 16 GT/s |
Operating system | Red Hat Enterprise Linux 8.3 (4.18.0-240.el8.x86_64) | Red Hat Enterprise Linux 7.8 |
Memory | DDR4 256 G (16 GB x 16) 3200 MT/s | |
BIOS/CPLD | 2.0.2 / 1.1.12 |
|
Interconnect | NVIDIA Mellanox HDR | NVIDIA Mellanox HDR 100 |
Table 2: Benchmark datasets used for GROMACS performance evaluation
Datasets | Details |
1536 K and 3072 K | |
1400 K and 3000 K | |
Prace – Lignocellulose | 3M |
The following information describes the performance evaluation for the processor stack listed in the Table 1.
Rome processors compared to Milan processors (GROMACS)
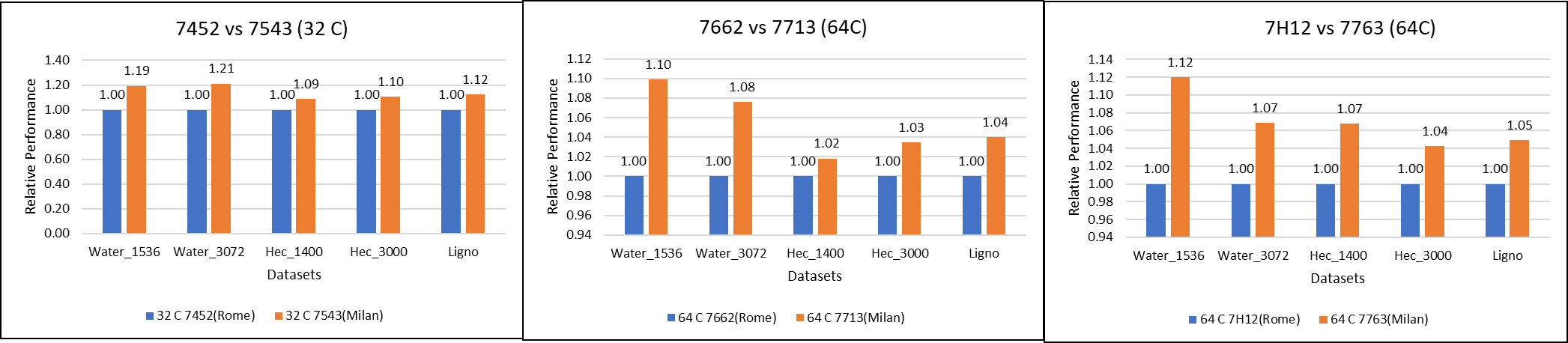 Figure 1: GROMACS performance comparison with AMD Rome processors
Figure 1: GROMACS performance comparison with AMD Rome processors
For performance benchmark comparisons, we selected Rome processors that are closest to their Milan counterparts in terms of hardware features such as cache size, TDP values, and Processor Base/Turbo Frequency, and marked the maximum value attained for Ns/day by each of the datasets mentioned in Table 2.
Figure 1 shows a 32C Milan processor has higher performance improvements (19 percent for water 1536, 21 percent for water 3072, and 10 to approximately 12 percent with HECBIO sim and lingo cellulose datasets) compared to a 32C Rome processor. This result is due to a higher processor speed and improved L3 cache, wherein more data can be accessed by each core.
Next, with the higher end processor we see only 10 percent gain with respect to the water dataset, as they are more memory intensive. Some percentage is added on due to improvement of frequency for the remaining datasets. Overall, the Milan processor results demonstrated a substantial performance improvement for GROMACS over Rome processors.
Milan processors comparison (32C processors compared to 64C processors)
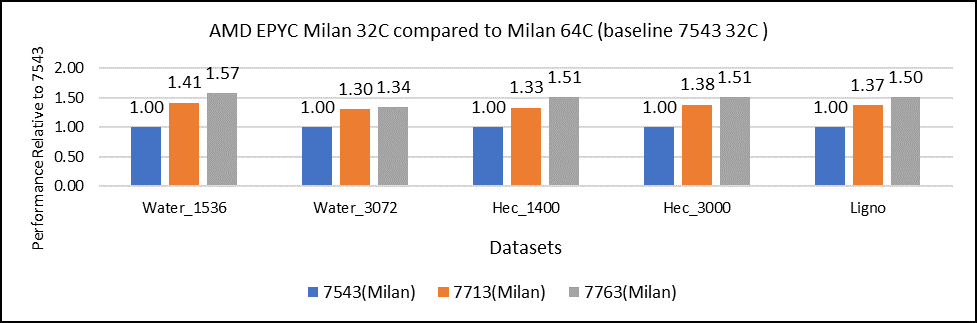
Figure 2: GROMACS performance with Milan processors
Figure 2 shows performance relative to the performance obtained on the 7543 processor. For instance, the performance of water 1536 is improved from the 32C processor to the 64 core (64C) processor from 41 percent (7713 processor) to 57 percent (7763 processor). The performance improvement is due to the increasing core counts and higher CPU core frequency performance improvement. We observed that GROMACS is frequency sensitive, but not to a great extent. Greater gains may be seen when running GROMACS across multiple ensembles runs or running dataset with higher number of atoms.
We recommend that you compare the price-to-performance ratio before choosing the processor based on the datasets with higher CPU core frequency, as the processors with a higher number of lower-frequency cores may provide better total performance.
Multi-node study with 7713 64C processors
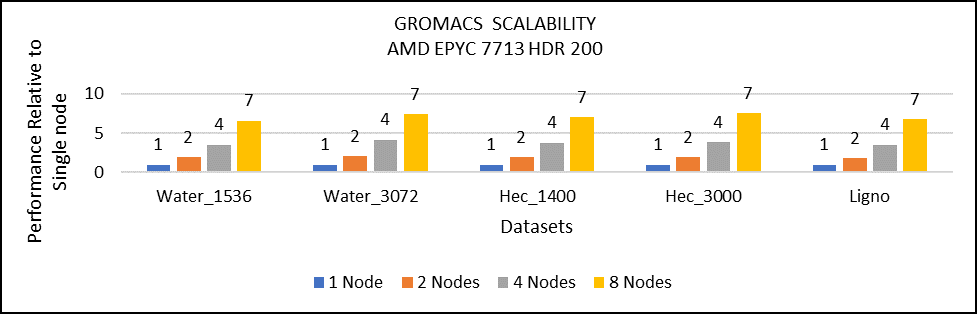
Figure 3: Multi-node study with 7713 64c SKUs
For multi-node tests, the test bed was configured with an NVIDIA Mellanox HDR interconnect running at 200 Gbps and each server included an AMD EPYC 7713 processor. We achieved the expected linear performance scalability for GROMACS of up to four nodes and across each of the datasets. All cores in each server were used while running the benchmarks. The performance increases are close to linear across all the dataset types as core count increases.
Conclusion
For the various datasets we evaluated, GROMACS exhibited strong scaling and was compute intensive. We recommend a processor with high core count for smaller datasets (water 1536, hec 1400); larger datasets (water 3072, ligno,HEC 3000) would benefit from memory per core. Configuring the best BIOS options is important to get the best performance out of the system.
For more information and updates, follow this blog site.
Related Blog Posts

Dell Reinforces its TPCx-AI Benchmark Leadership using the 16G PowerEdge R6625 Hardware Platform at SF1000
Wed, 12 Jul 2023 18:52:17 -0000
|Read Time: 0 minutes
Overview
On 06-13-2023, Dell Technologies published a TPCx-AI SF1000 result that was based on an 11 x Dell PowerEdge R6625 hardware platform powered by AMD Genoa processors. As of the publication date, Dell results held number one slots on the Top Performance and Price/Performance tables for TPCx-AI on SF3, SF100, SF300, and SF1000. These results reinforce Dell Technologies’ TPCx-AI benchmark leadership position; a statement to the great performance provided by its AI, ML, and DL solutions.
This blog presents the hardware platform that was tested, what was measured and what the results mean.
What TPCx-AI tests measure
TPCx-AI measures the end-to-end machine learning or data science platform using a diverse representative dataset scaling from 1 GB to 10 TB. The TPCx-AI benchmark assesses various aspects of AI training and inference performance, including data generation, model training, serving, scoring, and system scalability. The benchmark can be used across a wide range of different systems from edge to data center. It aims to provide a standardized and objective measure of AI performance across different platforms and configurations.
By using TPCx-AI, organizations and vendors can make informed decisions about the AI infrastructure that best suits their needs. The benchmark helps in understanding the system's capability to handle large-scale AI training workloads and can help optimize performance and resource allocation for AI tasks.
The TPCx-AI standard defines 10 use cases based on data science pipelines modeled on a retail business data center to evaluate the performance of artificial intelligence systems. The workload trains deep neural networks on large datasets using prominent machine learning frameworks such as TensorFlow. The benchmark measures:
- The total time taken to train a model for each use case to a specific level of accuracy
- The time taken for that model to be used for inference or serving
The blog, Interpreting the results of the TPCx-AI Benchmark, outlines the ten use cases, their data science models, and the benchmark phases.
System under test (SUT)
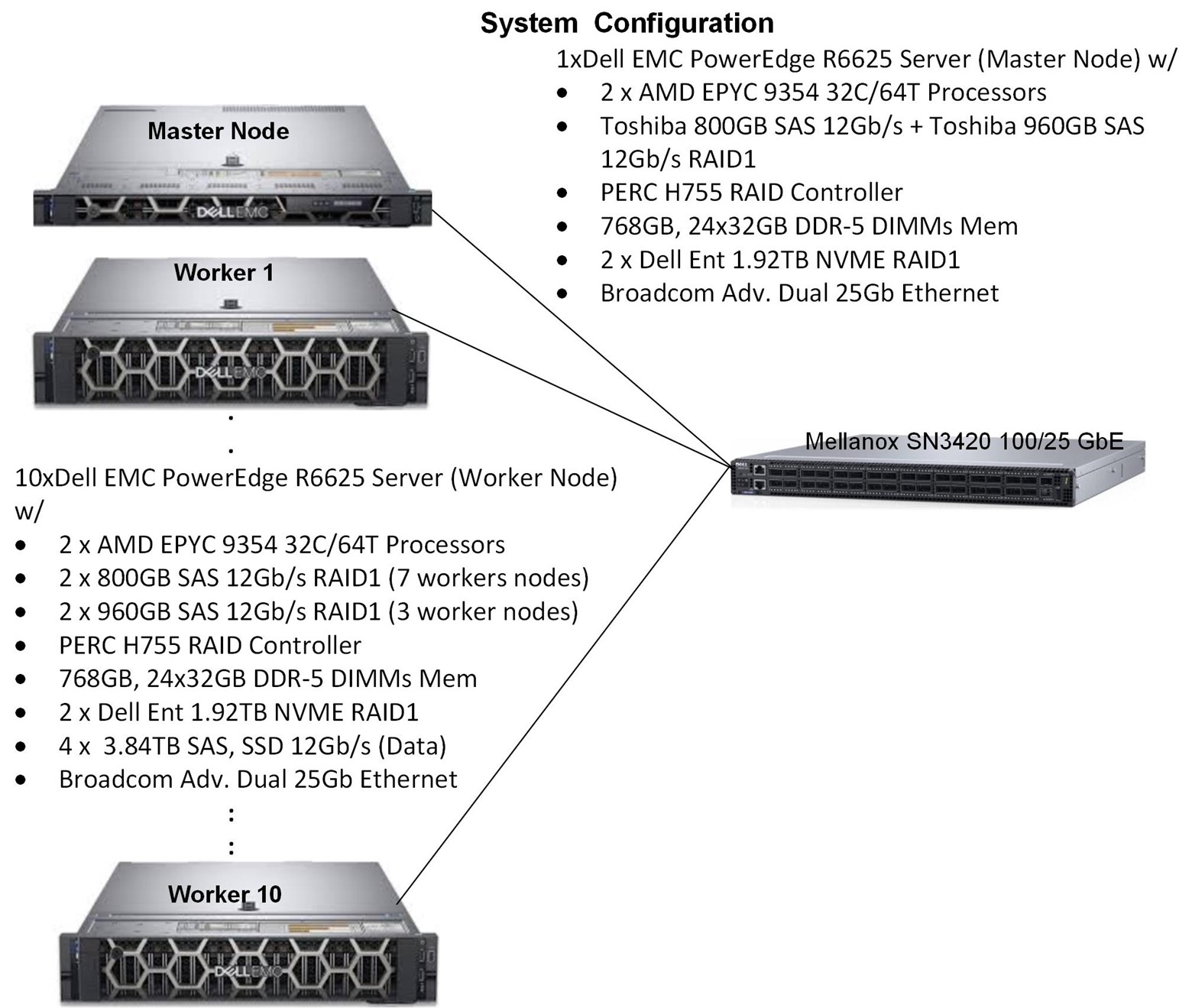
Figure 1 System Under Test (SUT).
Software versions
Table 1 Software versions
Software | Version |
Cloudera Data Platform (CDP) | 7.1.7 SP2 |
Hadoop | 3.1.1 |
HDFS | 3.1.1 |
YARN | 3.1.1 |
MR2 | 3.1.1 |
Spark | 2.4.7 |
ZooKeeper | 3.5.5 |
Java | 1.8.0 |
Python | 3.7.16 |
Red Hat Enterprise Linux | 8.7 (Master node) |
TPCx-AI Kit | 1.0.2 |
The result
Primary metrics
Table 2 Primary metric scores
Primary Metric | Score |
Performance (AIUCpm@1000) | 3,258.01 |
Price/Performance (USD/AIUCpm@100) | 267.96 |
Availability | June 13, 2023 |
The three primary metrics in Table 2 are required for all TPC results. The top ten results, based on performance or price/performance at a particular SF category, are displayed in the tables of the respective benchmark standard categorized by the metric and SF. To compare any results, all three metrics must be disclosed in the body of the message. The TPC does not allow comparing TPCx-AI results from different SF categories. The blog, Interpreting the results of the TPCx-AI Benchmark, goes into the details of how the performance and price/performance metrics are calculated. The availability date is the date all the priced line items (SKUs) are available to customers and must be within 185 days of the submission date. For the performance metric, the higher the score the better. For price/performance, the lower the better.
Other metrics
Table 3 Other metrics
Metric | Score |
Total system cost | $872,988 |
Framework | Cloudera SEL Data Platform Private Cloud Base Edition |
Operating system | Red Hat Enterprise Linux 8.6/8.7 |
Scale factor | 1,000 |
Physical storage divided by scale factor | 214.56 |
Scale factor divided by physical memory | 0.12 |
Main data redundancy mode | Replication 3, RAID 1 |
Number of servers | 11 |
Total processors, cores, and threads | 22/704/1,344 |
Number of streams | 4 |
The metrics in Table 3 are required to be reported and disclosed in the Full Disclosure Report (FDR) and Executive Summary (ES). Except for the total system cost, these other metrics are not used in the calculation of the primary metrics but provide additional information about the system that was tested. For instance, the total system cost is the total cost of ownership (TCO) for one year. The redundancy modes provide the data protection mechanisms that were used in the configuration as required by the benchmark standard. The number of streams refers to the number of concurrent serving tests during the Throughput phase.
Numerical quantities
Benchmark run times
Table 4 Benchmark run times
Benchmark run | Time |
Benchmark start | 06-07-2023 9:35:25 PM |
Benchmark end | 06-08-2023 3:20:10 AM |
Benchmark duration | 5:44:45.193 |
Benchmark phase times
Table 5 Benchmark phase metrics
Benchmark phase | Metric_name | Metric value |
Data Generation | DATAGEN | 2419.613 |
Data Loading | TLOAD | 927.45 |
Load Test | TLD | 927.45 |
Power Training | TPTT | 492.143 |
Power Serving 1 | TPST1 | 56.998 |
Power Serving 2 | TPST2 | 57.357 |
Power Serving | TPST | 57.357 |
Throughput | TTT | 43.934 |
AIUCpm@1000.0 | 3258.066 |
The seven benchmark phases and their metrics are explained in Interpreting the results of the TPCx-AI Benchmark, and are performed sequentially from data generation to throughput tests. In power training, models are generated and trained for each use case sequentially from UC1 to UC10. In power serving, the models obtained during the training phase are used to conduct the serving phase sequentially, one use case at a time. There are two power serving tests. The test that registers the longer time provides the TPST metric. The throughput phase runs multiple streams of serving tests concurrently. The more the number of streams, the more the system resources are taxed. Typically, the number of streams are increased until TTTn+1 > TTTn (where n+1 refers to the next throughput test). The duration of the longest running stream (TTPUT) is used to calculate the throughput test metric TTT.
Use case times and accuracy
Table 6 Use case times and accuracy
Use case | TRAINING | SERVING_1 | SERVING_2 | Throughput | Accuracy | Threshold |
1 | 523.703 | 51.215 | 49.736 | 56.083 | -1.00000 | -1.0 >= -1 |
2 | 1813.764 | 85.354 | 88.783 | 129.274 | 0.43830 | word_error rate <= 0.5 |
3 | 95.795 | 12.443 | 12.811 | 13.84 | 4.57451 | mean_squared_log_error <= 5.4 |
4 | 59.08 | 25.475 | 25.489 | 31.016 | 0.71189 | f1_score >= 0.65 |
5 | 943.023 | 76.289 | 78.351 | 91.615 | 0.03347 | mean_squared_log_error <= 5.4 <= 0.5 |
6 | 435.865 | 33.135 | 33.071 | 37.12 | 0.21355 | matthews_corrcoef >= 0.19 |
7 | 43.585 | 15.317 | 15.3 | 17.143 | 1.65306 | median_absolute_error <= 1.8 |
8 | 1940.283 | 338.579 | 341.811 | 372.418 | 0.74996 | accuracy_score >= 0.65 |
9 | 5448.735 | 703.291 | 699.631 | 745.458 | 1.00000 | accuracy_score >= 0.9 |
10 | 818.635 | 28.326 | 28.19 | 31.162 | 0.81691 | accuracy_score >= 0.7 |
Table 6 shows the use case run times (in seconds) for each benchmark phase and the accuracy of the model that was used. For instance, the RNN model that was generated and trained for UC2 had a word_error rate of 0.4383 which was less (better) than the threshold error_rate of 0.5. The XGBoost model trained for UC8 was 74.99% accurate which was above and better than the 65% minimum accuracy threshold requirement.
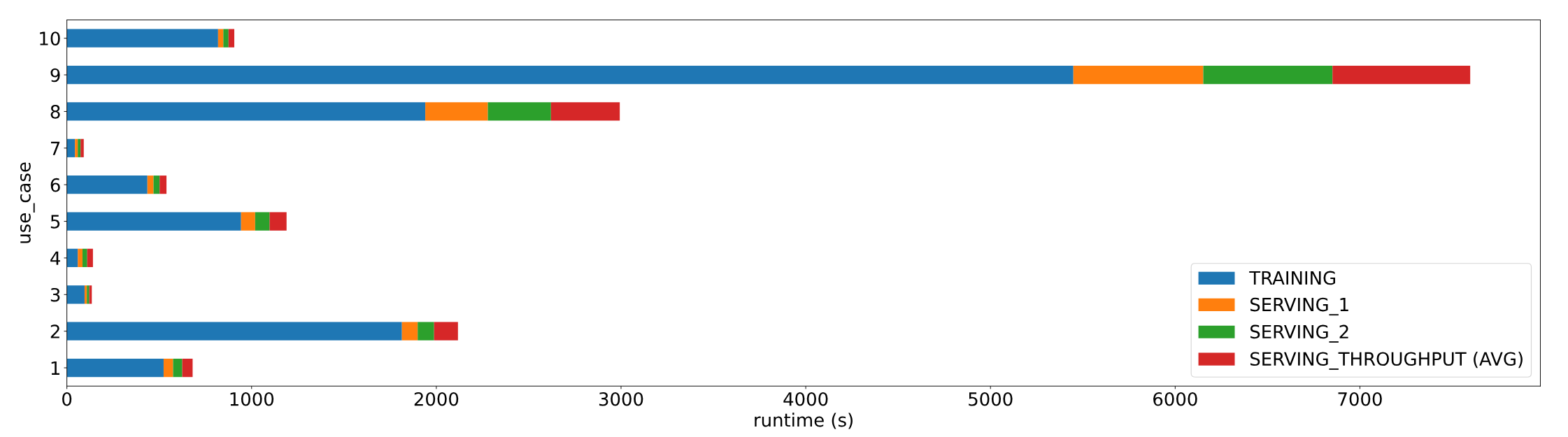
Figure 2 Use case time by benchmark phase
TPCx-AI SF1000 results tables
Table 7 displays the top TPCx-AI SF1000 tables as of the publication of this blog.
Table 7 SF1000 top performance table
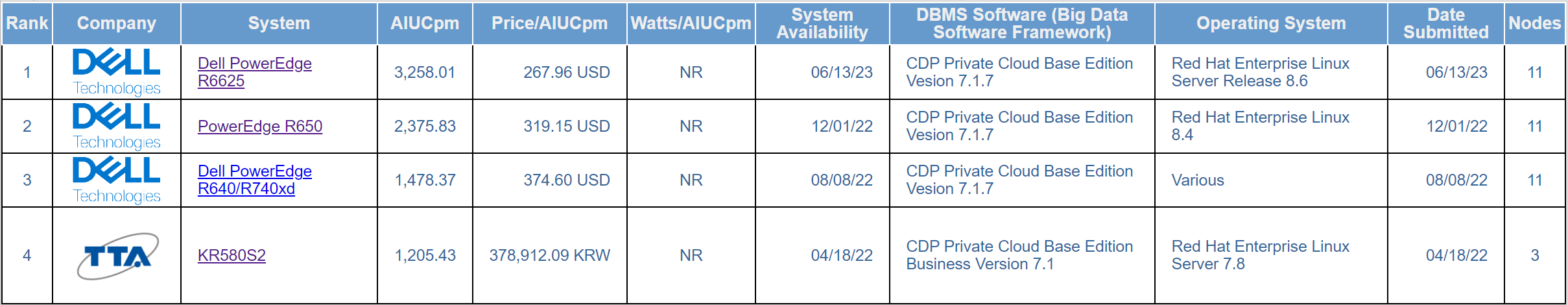
Table 8 Top price/performance table
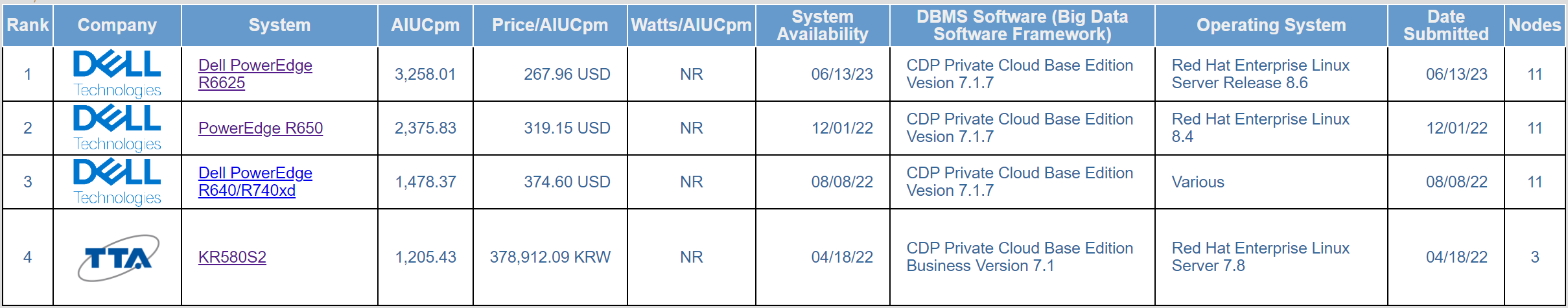
Table 7 and Table 8 are similar. Of the four published results at SF1000, Dell Technologies’ hardware platforms hold the number 1, number 2, and number 3 positions on both the performance and price/performance tables. The main difference between the three top results is the processor generations:
- The number 1 result used 4th generation AMD Genoa processors
- The number 2 result used 3rd generation Intel Ice Lake processors
- The number 3 result used 2nd generation Intel Cascade Lake processors
Key takeaways
- Dell dominates TPCx-AI top performance and price/performance tables at SF3, SF100, SF300, and SF1000.
- TPCx-AI performance improved greatly on newer generation Dell hardware platforms that have newer generation processors:
- There was a 60.71% performance improvement between hardware platforms powered by (14G) 2nd generation and (15G) 3rd generation processors.
- There was a 37.13% improvement between 3rd generation and (16G) 4th generation processors.
- TPCx-AI price/performance improved greatly between processor generations of the Dell 14G, 15G, and 16G hardware platforms:
- There was a 14.80% price/performance drop from hardware platforms powered by 2nd generation to 3rd generation processors.
- There was a 27.08% price/performance drop from 3rd generation to 4th generation processors.
- The form factor of the hardware platforms has reduced:
- The Dell 14G TPCx-AI SF1000 result used 2U servers
- The 15G and 16G results used 1U servers and scored better performance and price/performance
- Using NVMe data storage scored better price/performance metrics:
- The 14G result used hard drives
- The 15G and 16G results used more expensive NVMe data drives, and yet scored better price/performance metrics
Conclusion
This blog examined in detail the TPCx-AI performance result of the Dell 16G PE R6625 hardware platform. The result cemented Dell Technologies’ leadership positions on TPCx-AI performance and price/performance tables at SF1000, in addition to the leadership positions at SF3, SF100, and SF300. These results prove Dell Technologies’ leadership as a provider of high-performance AI, ML, and DL solutions based on verifiable performance data backed by a reputable, industry-standards performance consortium.
References
Nicholas Wakou, Nirmala Sundararajan; Interpreting the results of the TPCx-AI Benchmark; infohub.delltechnologies.com (February 2023).

HPC Application Performance on Dell PowerEdge R7525 Servers with NVIDIA A100 GPGPUs
Tue, 24 Nov 2020 17:49:03 -0000
|Read Time: 0 minutes
Overview
The Dell PowerEdge R7525 server powered with 2nd Gen AMD EPYC processors was released as part of the Dell server portfolio. It is a 2U form factor rack-mountable server that is designed for HPC workloads. Dell Technologies recently added support for NVIDIA A100 GPGPUs to the PowerEdge R7525 server, which supports up to three PCIe-based dual-width NVIDIA GPGPUs. This blog describes the single-node performance of selected HPC applications with both one- and two-NVIDIA A100 PCIe GPGPUs.
The NVIDIA Ampere A100 accelerator is one of the most advanced accelerators available in the market, supporting two form factors:
- PCIe version
- Mezzanine SXM4 version
The PowerEdge R7525 server supports only the PCIe version of the NVIDIA A100 accelerator.
The following table compares the NVIDIA A100 GPGPU with the NVIDIA V100S GPGPU:
NVIDIA A100 GPGPU | NVIDIA V100S GPGPU | |||
Form factor | SXM4 | PCIe Gen4 | SXM2 | PCIe Gen3 |
GPU architecture | Ampere | Volta | ||
Memory size | 40 GB | 40 GB | 32 GB | 32 GB |
CUDA cores | 6912 | 5120 | ||
Base clock | 1095 MHz | 765 MHz | 1290 MHz | 1245 MHz |
Boost clock | 1410 MHz | 1530 MHz | 1597 MHz | |
Memory clock | 1215 MHz | 877 MHz | 1107 MHz | |
MIG support | Yes | No | ||
Peak memory bandwidth | Up to 1555 GB/s | Up to 900 GB/s | Up to 1134 GB/s | |
Total board power | 400 W | 250 W | 300 W | 250 W |
The NVIDIA A100 GPGPU brings innovations and features for HPC applications such as the following:
- Multi-Instance GPU (MIG)—The NVIDIA A100 GPGPU can be converted into as many as seven GPU instances, which are fully isolated at the hardware level, each using their own high-bandwidth memory and cores.
- HBM2—The NVIDIA A100 GPGPU comes with 40 GB of high-bandwidth memory (HBM2) and delivers bandwidth up to 1555 GB/s. Memory bandwidth with the NVIDIA A100 GPGPU is 1.7 times higher than with the previous generation of GPUs.
Server configuration
The following table shows the PowerEdge R7525 server configuration that we used for this blog:
Server | PowerEdge R7525 |
Processor | 2nd Gen AMD EPYC 7502, 32C, 2.5Ghz |
Memory | 512 GB (16 x 32 GB @3200MT/s) |
GPGPUs | Either of the following: 2 x NVIDIA A100 PCIe 40 GB 2 x NVIDIA V100S PCIe 32 GB |
Logical processors | Disabled |
Operating system | CentOS Linux release 8.1 (4.18.0-147.el8.x86_64) |
CUDA | 11.0 (Driver version - 450.51.05) |
gcc | 9.2.0 |
MPI | OpenMPI-3.0 |
HPL | hpl_cuda_11.0_ompi-4.0_ampere_volta_8-7-20 |
HPCG | xhpcg-3.1_cuda_11_ompi-3.1 |
GROMACS | v2020.4 |
Benchmark results
The following sections provide our benchmarks results with observations.
High-Performance Linpack benchmark
High Performance Linpack (HPL) is a standard HPC system benchmark. This benchmark measures the compute power of the entire cluster or server. For this study, we used HPL compiled with NVIDIA libraries.
The following figure shows the HPL performance comparison for the PowerEdge R7525 server with either NVIDIA A100 or NVIDIA V100S GPGPUs:
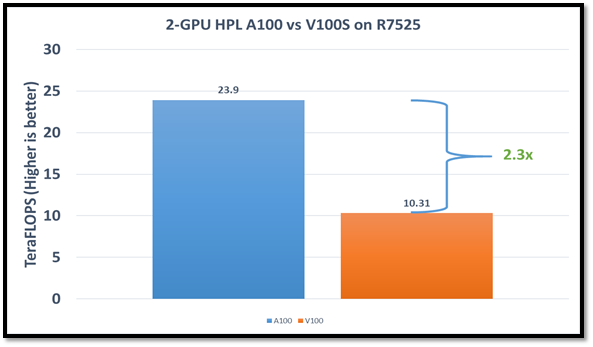
Figure1: HPL performance on the PowerEdge R7525 server with the NVIDIA A100 GPGPU compared to the NVIDIA V100SGPGPU
The problem size (N) is larger for the NVIDIA A100 GPGPU due to the larger capacity of GPU memory. We adjusted the block size (NB) used with the:
- NVIDIA A100 GPGPU to 288
- NVIDIA V100S GPGPU to 384
The AMD EPYC processors provide options for multiple NUMA combinations. We found that the best value of 4 NUMA per socket (NPS=4), with NUMA per socket 1 and 2 lower the performance by 10 percent and 5 percent respectively. In a single PowerEdge R7525 node, the NVIDIA A100 GPGPU delivers 12 TF per card using this configuration without an NVLINK bridge. The PowerEdge R7525 server with two NVIDIA A100 GPGPUs delivers 2.3 times higher HPL performance compared to the NVIDIA V100S GPGPU configuration. This performance improvement is credited to the new double-precision Tensor Cores that accelerate FP64 math.
The following figure shows power consumption of the server while running HPL on the NVIDIA A100 GPGPU in a time series. Power consumption was measured with an iDRAC. The server reached 1038 Watts at peak due to a higher GFLOPS number.
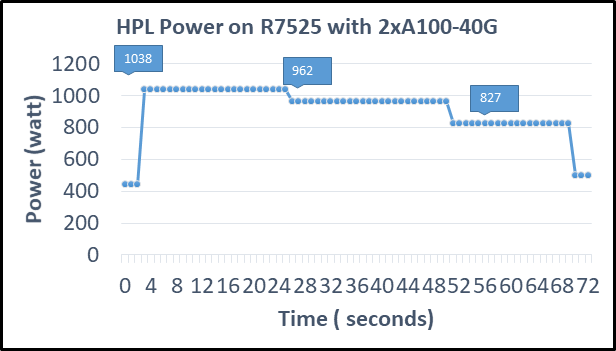
Figure2: Power consumption while running HPL
High Performance Conjugate Gradient benchmark
The High Performance Conjugate Gradient (HPCG) benchmark is based on a conjugate gradient solver, in which the preconditioner is a three-level hierarchical multigrid method using the Gauss-Seidel method.
As shown in the following figure, HPCG performs at a rate 70 percent higher with the NVIDIA A100 GPGPU due to higher memory bandwidth:
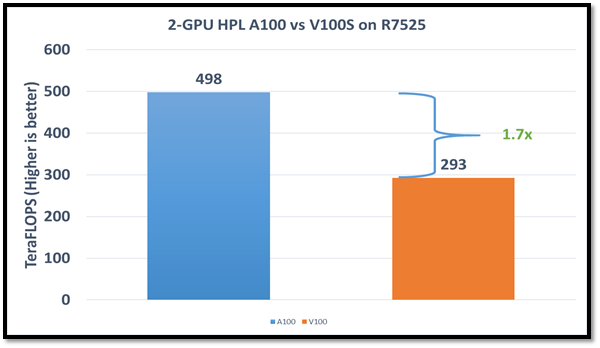
Figure 3: HPCG performance comparison
Due to different memory size, the problem size used to obtain the best performance on the NVIDIA A100 GPGPU was 512 x 512 x 288 and on the NVIDIA V100S GPGPU was 256 x 256 x 256. For this blog, we used NUMA per socket (NPS)=4 and we obtained results without an NVLINK bridge. These results show that applications such as HPCG, which fits into GPU memory, can take full advantage of GPU memory and benefit from the higher memory bandwidth of the NVIDIA A100 GPGPU.
GROMACS
In addition to these two basic HPC benchmarks (HPL and HPCG), we also tested GROMACS, an HPC application. We compiled GROMACS 2020.4 with the CUDA compilers and OPENMPI, as shown in the following table:
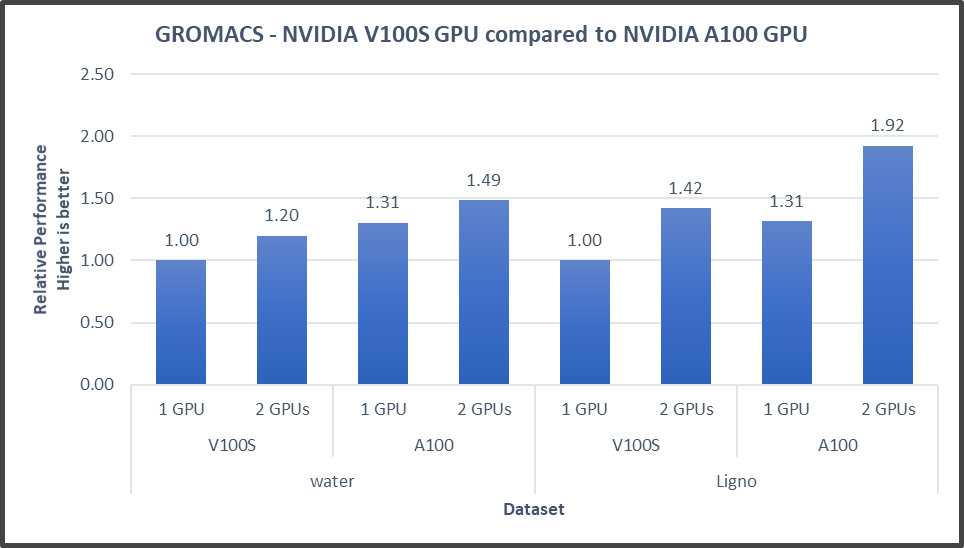
Figure4: GROMACS performance with NVIDIA GPGPUs on the PowerEdge R7525 server
The GROMACS build included thread MPI (built in with the GROMACS package). All performance numbers were captured from the output “ns/day.” We evaluated multiple MPI ranks, separate PME ranks, and different nstlist values to achieve the best performance. In addition, we used settings with the best environment variables for GROMACS at runtime. Choosing the right combination of variables avoided expensive data transfer and led to significantly better performance for these datasets.
GROMACS performance was based on a comparative analysis between NVIDIA V100S and NVIDIA A100 GPGPUs. Excerpts from our single-node multi-GPU analysis for two datasets showed a performance improvement of approximately 30 percent with the NVIDIA A100 GPGPU. This result is due to improved memory bandwidth of the NVIDIA A100 GPGPU. (For information about how the GROMACS code design enables lower memory transfer overhead, see Developer Blog: Creating Faster Molecular Dynamics Simulations with GROMACS 2020.)
Conclusion
The Dell PowerEdge R7525 server equipped with NVIDIA A100 GPGPUs shows exceptional performance improvements over servers equipped with previous versions of NVIDIA GPGPUs for applications such as HPL, HPCG, and GROMACS. These performance improvements for memory-bound applications such as HPCG and GROMACS can take advantage of higher memory bandwidth available with NVIDIA A100 GPGPUs.