

Tuxedo Pipeline Performance on Dell EMC PowerEdge C6520
Tue, 22 Jun 2021 15:00:00 -0000
|Read Time: 0 minutes
Overview
Gene expression analysis is as important as identifying Single Nucleotide Polymorphism (SNP), insertion/deletion (indel), or chromosomal restructuring. Ultimately, all physiological and biochemical events depend on the final gene expression product, proteins. Although most mammals have an additional controlling layer before protein expression, knowing how many transcripts exist in a system helps to characterize the biochemical status of a cell. Ideally, technology should enable us to quantify all proteins in a cell, which would advance the progress of Life Science significantly; however, we are far from achieving this.
In this blog, we report the test results of one popular RNA-Seq data analysis pipeline known as the Tuxedo pipeline. The Tuxedo pipeline suite offers a set of tools for analyzing a variety of RNA-Seq data, including short-read mapping, identification of splice junctions, transcript and isoform detection, differential expression, visualizations, and quality control metrics. The tested workflow is a differentially expressed gene (DEG) analysis, and the detailed steps in the pipeline are shown in Figure 1.
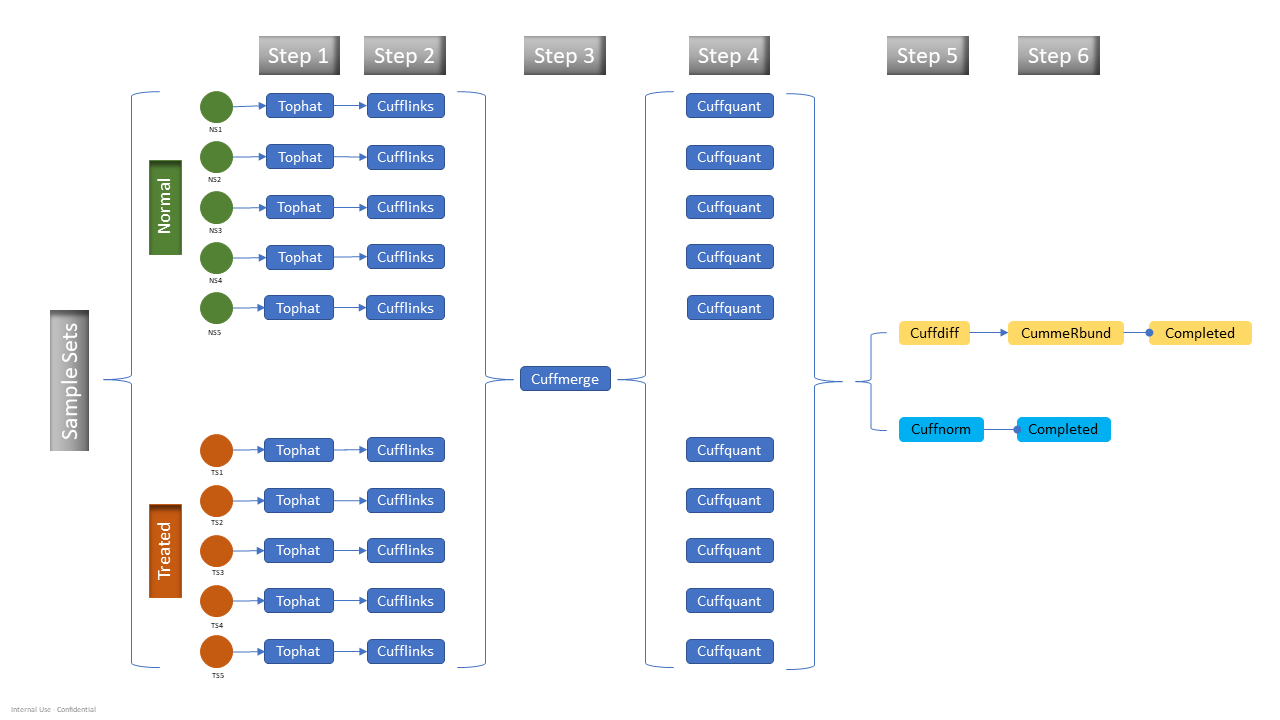
Figure 1: Updated Tuxedo Pipeline with Cuffquant Step
In this study, the performances of single nodes with 3rd Gen Intel® Xeon® Scalable Processors (Codename Ice Lake) and 2nd Generation Intel® Xeon® Scalable Processors (Codename Cascade Lake) on Dell EMC PowerEdge R6520 (liquid-cooled) servers and C6420 (air-cooled) servers were compared. The configurations of the test systems are summarized in Table 1.
Table 1: Tested compute node configuration
Dell EMC PowerEdge C6520 Liquid Cooled | |
CPU | Tested 3rd Gen Intel® Xeon® Scalable Processors: 2 x Intel® Xeon® Platinum 8358, 32 Cores, 2.60 GHz – 3.40 GHz Base-Boost, TDP 250W 2 x Intel® Xeon® Platinum 8352Y, 32 Cores, 2.20 GHz – 3.40 GHz Base-Boost, TDP 205W
Tested 2nd Generation Intel® Xeon® Scalable Processors: 2 x Intel® Xeon® Gold 6248, 20 Cores, 2.50 GHz – 3.90 GHz Base-Boost, TDP 150W on Dell EMC PowerEdge C6420 Air Cooled |
RAM | DDR4 512 GB (16 x 32 GB) 3200 MT/s |
Operating system | RHEL 8.3 (4.18.0-240.el8.x86_64) |
Interconnect | Mellanox InfiniBand HDR100 |
File system | Dell EMC Ready Solutions for HPC BeeGFS High Capacity Storage |
BIOS system profile | Performance Optimized |
Logical processor | Disabled |
Virtualization technology | Disabled |
tophat | 2.1.1 |
bowtie2 | 2.2.5 |
R | 3.6 |
bioconductor-cummerbund | 2.26.0 |
A performance study of the RNA-Seq pipeline is not trivial because the nature of the workflow requires input files that are non-identical but similar in size. Hence, 185 RNA-Seq paired-end read data are collected from a public data repository. All the read data files contain around 25 Million Fragments (MF) and have similar read lengths. The samples for a test are randomly selected from the pool of the 185 paired-end read files. Although these test data will not have any biological meaning, certainly these data with a high level of noise will put the tests in the worst-case scenario.
Performance Evaluation
Throughput Test – Single pipeline with more than two samples, biological and technical duplicates
Typical RNA-Seq studies consist of multiple samples, sometimes hundreds of different samples, for example, normal versus disease or untreated versus treated samples. These samples tend to have a high level of noise for biological reasons; hence, the analysis requires vigorous data preprocessing procedures.
We tested various numbers of samples (all different RNA-Seq data selected from 185 paired-end reads data sets) to see how much data can be processed by a single node. Typically, when the number of samples increases, the runtime of the Tuxedo Pipeline increases as shown in Figure 2. Ice Lake CPUs show improved overall runtime of 10% and more compared to Cascade Lake 6248 CPUs.
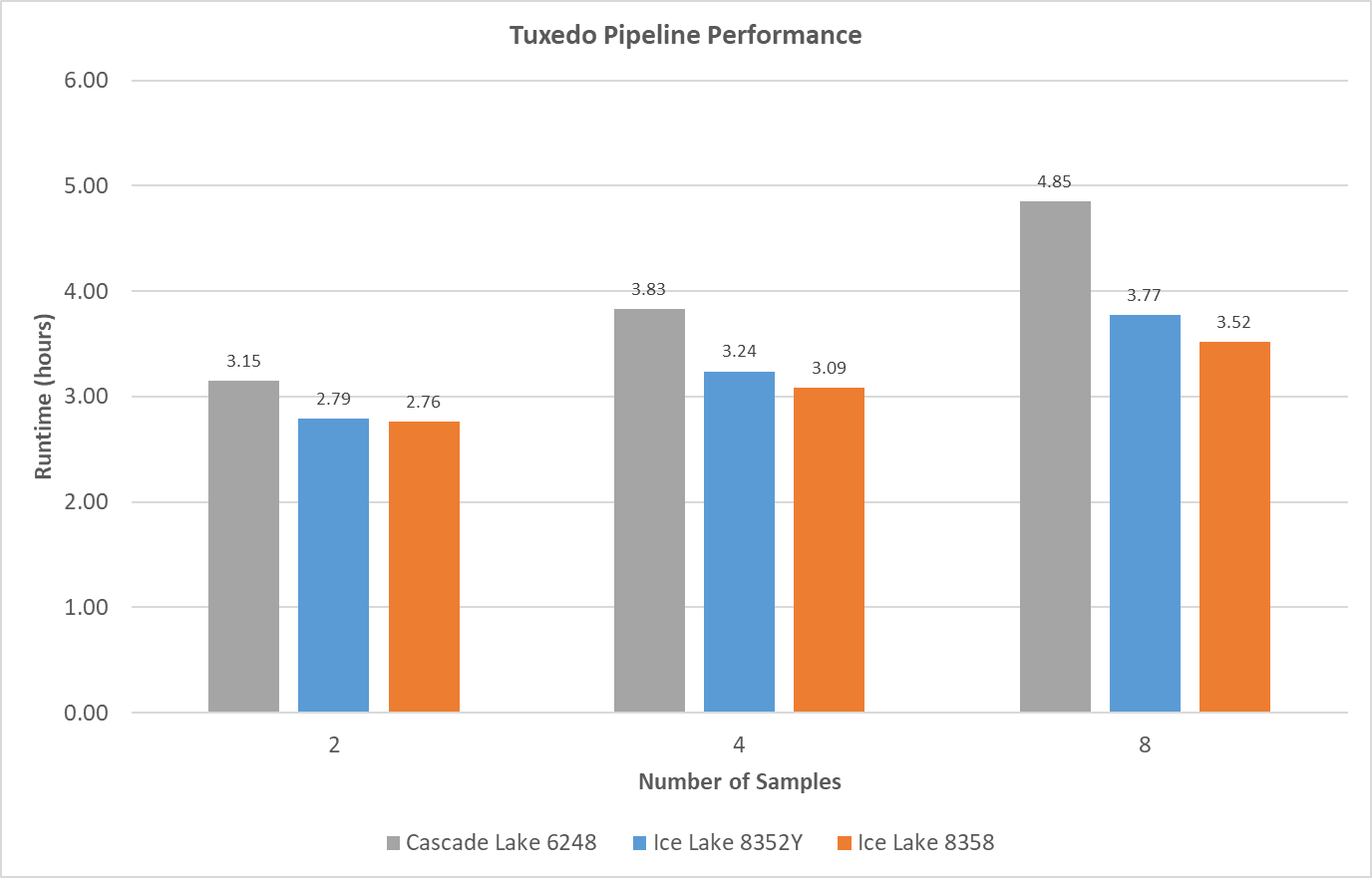
Figure 2: Total runtime comparisons from various number of samples with a single compute node
Conclusion
Many additional tests are still required to obtain a better insight from Intel Ice Lake processors for the NGS data analysis area. Unfortunately, we could not push our tests over 8 samples due to the storage limitation. However, there seems to be plenty of room for a higher throughput processing of more than 8 samples together.


