Home > Workload Solutions > High Performance Computing > White Papers > Dell Validated Design for Health Care and Life Sciences with Dell PowerEdge C6520 and XE8545 > Clara Parabricks secondary analysis pipeline
Clara Parabricks secondary analysis pipeline
-
Analysis performed on NGS data is often described as a pipeline. A pipeline is a defined workflow that consists of a collection of methods or operations where the output of one operation becomes the input for the next operation. Four critical operations—mapping, alignment, preprocessing, and variant calling—make up most secondary analysis WGS pipelines. Clara Parabricks is a software suite for genomic analysis methods designed to take advantage of GPU acceleration. Many of the Clara Parabricks methods are functionally equivalent to existing open-source methods, often generating >99.9% concordance. Clara Parabricks operations are stitched together to create a secondary analysis pipeline best matched to the requirements for the sequencing application of interest such as germline or somatic analysis. Clara Parabricks is available as either a Docker® or Singularity container and uses various server GPU resources. Figure 9 highlights the Clara Parabricks v4.0.0-1 application suite.

Figure 9. Parabricks application suite
Calling genetic variants present in an individual genome relies on millions to billions of short, error-prone sequence reads. Despite over a decade of effort by thousands of dedicated researchers, the hand-crafted and parameterized statistical models used for variant calling produce thousands of errors and missed variants in each genome (Poplin, 2016). Many groups run consensus variant calling pipelines using multiple variant calling method to minimize the likelihood of missing a variant. Clara Parabricks contains multiple variant callers to enable this approach. For this study, the germline pipeline was used, and the steps are listed in Figure 10.
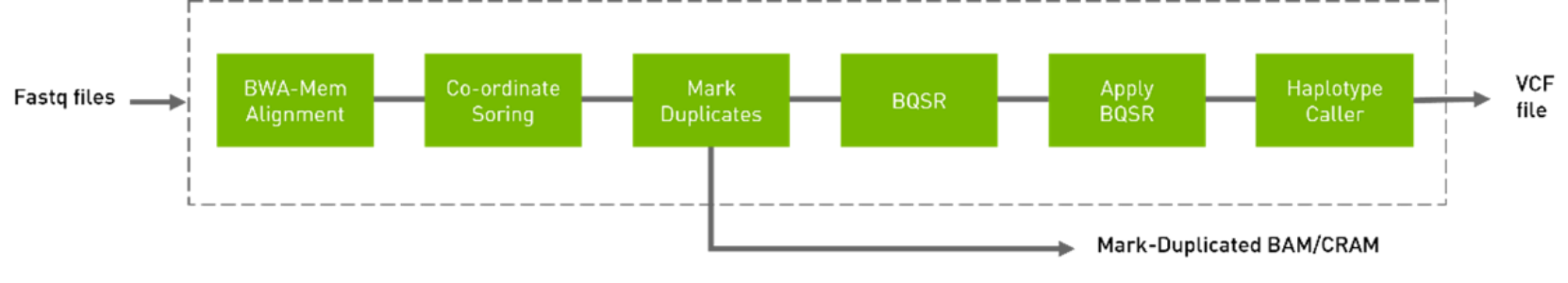
Figure 10. Germline pipeline
DeepVariant, a variant calling method developed by Google®, applies a deep convolutional neural network and has been shown to outperform expert driven statistical methods. However, calling variants for a 30x human genome and writing the variants out to a gVCF file takes approximately four hours and requires at least 1,024 compute cores. The NVIDIA Clara Parabricks GPU accelerated version of DeepVariant runs in less than 20 minutes for a 30X genome. The fast analysis time enables using DeepVariant alone or combined with other germline callers like GATK HaplotypeCaller, while minimizing the potential of creating a secondary analysis backlog. DeepVariant v1.4 is now available in the Clara Parabricks collection on NGC. It brings significant improvements to how genomic researchers and bioinformaticians deploy and scale genome sequencing analysis pipelines. The first of these releases is for DeepVariant v1.4. This latest version of DeepVariant increases accuracy across multiple genomics sequencers.
An additional read insert size feature for Illumina whole genome and whole exome models reduces errors by 4-10%. It uses direct phasing for more accurate variant calling in PacBio sequencing runs. You can now perform the high-accuracy process of phased variant calling for PacBio data directly in DeepVariant, with pipelines such as DeepVariant-WhatsHap-DeepVariant or PEPPER-Margin-DeepVariant.
